Association of Aberrant Promoter Methylation Changes in the Suppressor of Cytokine Signaling 3 (SOCS3) Gene with Susceptibility to Crohn's Disease
-
Sanati, Golshid
-
Department of Immunology, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
-
Jafari, Davood
-
Department of Immunology, School of Medicine, Zanjan University of Medical Sciences, Zanjan, Iran
-
Immunogenetics Research Network (IgReN), Universal Scientific Education and Research Network (USERN), Zanjan, Iran
-
Noruzinia, Mehrdad
-
Department of Medical Genetics, Faculty of Medical Sciences, Tarbiat Modares University, Tehran, Iran
-
Ebrahimi-Daryani, Naser
-
Department of Internal Medicine, Division of Gastroenterology, Imam Khomeini Hospital, Tehran University of Medical Sciences, Tehran, Iran
-
Ahmadvand, Mohammad
-
Department of Medical Genetics, Faculty of Medical Sciences, Tarbiat Modares University, Tehran, Iran
-
Teimourian, Shahram
-
Department of Genetics, School of Medicine, Iran University of Medical Sciences, Tehran, Iran
-
 Rezaei, Nima
Children's Medical Center Hospital, Tehran, Iran, rezaei_nima@tums.ac.ir
Rezaei, Nima
Children's Medical Center Hospital, Tehran, Iran, rezaei_nima@tums.ac.ir
-
Department of Immunology, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
-
Research Center for Immunodeficiencies, Children's Medical Center, Tehran University of Medical Sciences, Tehran, Iran
-
Network of Immunity in Infection, Malignancy and Autoimmunity (NIIMA), Universal Scientific Education and Research Network (USERN), Boston, USA
Abstract: Background: Growing evidence supports that changes in the methylation state of Inflammatory Bowel Disease (IBD)-associated genes could significantly alter levels of gene expression, potentially contributing to disease onset and progression. We supposed that alterations in DNA methylation status at promoter region within the suppressor of cytokine signaling 3 (SOCS3) gene in intestinal tissues may be involved in the susceptibility to Crohn's Disease (CD).
Methods: DNA methylation status in the promoter region of the human SOCS3 gene of intestinal tissues from 15 patients with CD and 15 age- and sex-matched healthy controls were profiled using the real-time Quantitative Multiplex Methylation Specific PCR (QM-MSP) assay.
Results: Based on methylation assay data profiling, we found that patients with CD showed a higher degree of methylation of the SOCS3 gene promoter region than did the healthy controls (unmethylated DNA in CD vs. healthy controls; 0.00048±0.0011 vs. 0.07±0.142, p<0.000).
Conclusion: The data presented here demonstrate that aberrant methylation of the CpG islands within promoter regions of SOCS3 gene in colonic mucosa of CD was associated with mucosal inflammatory status, providing insights into the involvement of methylation could contribute to the initiation of the inflammatory process and development of CD.
Introduction :
Crohn's Disease (CD) is a major subtype of Inflammatory Bowel Disease (IBD) and could potentially affect whole Gastrointestinal (GI) tract 1. CD is histopathologically characterized by the presence of persistent inflammation of the terminal ileum and colon dominates 2,3. CD appears to be multifactorial and polygenic and the precise etiology of the disease is not clear. However, it is assumed that the onset and perpetuation of the disease is caused by a complex inter-play between susceptibility genes, the immune system and environmental factors 3,4.
It is becoming apparent that epigenetic factors can have a significant contribution in the pathogenesis of disease 5. Epigenetic can be defined as inherited changes in gene expression that is not involved in DNA sequencing itself and is influenced by environment 6. Genome Wide Association Studies (GWAS) have reported more than 160 gene loci related to IBD, but only 13.6% of CD patients and 7.5% of ulcerative colitis patients are accounted for this IBD-associated genetic variants so the role of non-genetic factors such as epigenetic regulations have been bolded recently 7. Emerg-ing evidence suggests that epigenetic modifications, including changes in the methylation state of IBD-associated genes could significantly alter levels of gene expression, potentially contributing to disease onset and progression 8.
Jack/stat pathway is cytokine-modulated intracellular signaling pathway which can regulate the immune system development and hematopoiesis potentially through inflammatory cytokines such as Tumor Necrosis Factor (TNF)-α, Interferon (IFN)-γ and Interleukin (IL)-6 2,9. STAT family of transcription factors including STAT3 have been demonstrated to have a pivotal role in transmitting inflammatory cytokine signals to the nucleus 10. The suppressors of cytokine signaling (SOCS) 3 protein as a member of SOCS family is the key physiological regulators of cytokine-mediated activation of STAT3 by inhibiting the catalytic activity of JACKs 11. In the recent decades, SOCS3 has been one of the major interesting research subjects due to changes in the expression of SOCS3 contribute to the development of a variety of autoimmune diseases including IBDs 2,9.
It has been shown that in IBD patients, SOCS3 expression is considerably decreased in comparison with healthy volunteers 12-14, however up to our knowledge there is no study that investigate the SOCS3 promoter region methylation as a fundamental epigenetic mechanism. To appreciate the role of DNA methylation as a mechanism underlying differences in SOCS3 expression level and consequently in the regulation of the protein, we checked promoter methylation status. The primary goal of this study was to determine the alterations in DNA methylation status at promoter region within SOCS3 gene in intestinal tissues to clarify whether it is hyper or hypo-methylated in CD in comparison to normal population.
Materials and Methods :
Patients and tissue samples
We studied extracted DNA from colon biopsy specimens of 15 CD patients (8 females, 7 males), who un-derwent total proctocolectomy at the gastroenterology clinics of Kasra and Laleh Hospitals in Tehran, Iran, between May 2018 and July 2019. Colonic biopsies from 15 age- and sex-matched patients (8 females, 7 males) without abnormalities at colonoscopy and histories of gastrointestinal disease served as healthy controls. Diagnosis of CD was determined according to the usual clinical criteria, endoscopic and pathohistological criteria. The tissue samples were frozen at -80°C immediately. The study protocol was approved by the Committees on the Ethics of Human Research of Tehran University of Medical Sciences.
DNA isolation and Bisulfite conversion system
Genomic DNA was isolated from frozen colon tissues using the High Pure PCR Template Preparation Kit (Roche) according to the manufacturer’s protocol. DNA concentration and quality were measured using the NanoDrop ND-1000 spectrophotometer. Genomic DNA was modified by sodium bisulfite to convert unmethylated cytosines to uracil using the available MethylEdge™ Bisulfite Conversion System (Promega, Madison, WI) as per manufacturer’s instructions. Bisulfite-modified DNA specimens were aliquoted and stored at -20°C.
Methylation-specific PCR
The SOCS3 promoter and gene sequences, obtained from the GenBank database of National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm. nih.gov), were used to design primers for methylation studies. The MethySYBR procedure requires two sequential steps of PCR reactions. MethySYBR primers for SOCS3 gene were designed, including external for-ward primer (EXT-F1: 5'-GTAGGGAGGTGACGAG GTAGG-3'), external reverse primer (EXT-R: 5'- GGT TGTTCGGGGTTATTTTGT -3'), contain no CpG di-nucleotides. The external primers was used in the PCR reaction amplified total DNA templates of the target gene regardless of their methylation status. This served as a reference control to normalize the amount of methylated target alleles detected by the methylation specific primer pair between samples. The initial amplification of PCR reaction was performed in 25 µl of PCR reaction mixture composed of 1 µl bisulphite-treated genomic DNA, SOCS3 external primers 1 µl. The DNA was denatured at 95°C for 5 min, followed by 30 cycles at 94°C for 30 s, 56°C for 30 s, and 72°C for 30 s, with a final extension at 72°C for 5 min.
For designing the nested methylation-specific primers, three CpG sites of SOCS3 promoter were targeted in the primer sequence. The methylation profile of the promoter CpG islands were determined based on UCSC database. Methylation-Specific PCR (MSP)-specific primers were designed by Methprimer used to amplify methylated and unmethylated DNA sequences as appropriate. Methyl forward primer (FM: 5'-GGA GATTTTAGGTTTTCGGAATATTTC-3'), Methyl re-verse primer (RM: 5'-CCCCCGAAACTACCTAAAC GCCG-3'). The methylation status of bisulfite-con-verted DNA was determined using a fluorescencebased, real-time polymerase chain reaction (Real time-PCR) assay. For the second-step real-time PCR reactions, 1 µl of the diluted PCR product was used in each PCR reaction. The bisulfite treated DNA was PCR amplified in a 10 μl reaction volume containing containing 5 μl SYBR® Green Master Mix (Ampliqon, Denmark), 0.25 µl of each of the methylated primers, 3.5 µl DDW and 1 μl of bisulfite-treated DNA. Quantitative MSP was performed with a 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA).
No untreated template controls were included in each run as negative controls.
For data presentation, fully converted methylated human DNA was included in each run to serve as the 100% methylated reference for calculating the relative methylation percentages of DNA samples based on the relative 2-ΔΔCT quantitation approach.
Statistical analysis
DNA methylation data were expressed as mean±SD. The possible statistical differences in CpG island methylation levels between two groups were determined using the Mann-Whitney test. Chi-square statistic was used to discover if there is a relationship between two categorical variables. To calculate the mea-sure of association, the Odds Ratio (OR) and 95% confidence interval were considered. All p-values were 2-tailed and considered significant when less than 0.05. All statistical calculations were made using SPSS 21.0 version.
Results :
Because some genes are shown to be methylated with aging and sex, age- and sex-matched healthy controls were studied. The study consisted of 15 CD patients, with a mean age of 46±17.2 years (range: 17-71 years), and included 7 (46.7%) males and 8 (53.3%) females. Furthermore, fifteen were healthy controls (8 females, 7 males), with a mean age of 46±15.38 years (range: 28-78 years).
The use of SYBR green dye during quantitative PCR enables melting curve analysis of target amplicons to determine the methylation status of CpG sites on SOCS3 gene. The resulting melting curve for SOCS3 gene is shown in figure 1.
The promoter methylation status of SOCS3 gene in colonic mucosa specimens of all 15 CD patients and 15 paired healthy controls were compared (Figure 2). Methylation assay data profiling showed evidence of differential promoter methylation levels between patients with CD and healthy controls in which all three regions of the SOCS3 gene were more highly methylated in the active inflamed mucosa (unmethylated DNA= 0.00048±0.0011) than in mucosa of healthy controls (unmethylated DNA=0.07±0.142) (p<0.000, Figure 2).
Qualitative methylation data analysis of each individual showed a significant difference in the frequency of promoter methylation status between rectal inflammatory mucosa (12/15, 80%), and paired normal terminal ileum (2/15, 13.3%). The relative risk of disease indicated that individuals who had an aberrant promoter methylation profiling, were at high risk of the development of CD (p=0.009, OR=26.00, CI=22.85-337.72).
Discussion :
Chronic inflammation including IBD is tightly related to increased levels of DNA methylation 15,16. Although there have been studies reporting increased DNA methylation in colonic mucosa of IBD disease, the most recently published evidence in this field show the association between patterns of DNA methylation of colon mucosa and UC-associated dysplasia or carcinoma incidence without considering the mucosal inflammatory status 15. In the present study, we have examined and compared the alterations in CpG island methylation status at the SOCS3 gene promoter region in colorectal tissue specimens on the disease susceptibility. Our data demonstrated that methylation levels of SOCS3 gene-associated CpG islands were substantially higher in colorectal mucosa of patients with CD than in mucosa from healthy controls.
Despite the importance of the currently known genetic predispositions of multiple genes in the pathogenesis of IBD, an increasing number of observations reveal that abnormal epigenetic changes are considered as or even more important for IBD pathogenesis than genetic predisposition. Twin studies reveal that epigenetic modifications play an important etiologic role so that monozygotic twin on colectomy samples, one of who had CD methylation differences were observed in several genes that reveals the importance of the role of epigenetics. This observation suggests that prenatal developmental programming of colonic mucosal and immunological maturation imprints/programs postnatal microbiota shifts in the intestinal mucosa DNA methylation at different sites in a gene has quite different outcome in transcriptional activity 2,17.
Cytokines are secreted proteins central for coordination of the initiation, maintenance, and termination of all types of immune responses, including host responses to infection, inflammation. A tight control of cytokine release and of responses to cytokines is required for the defense against infections, the prevention of infection-associated immunopathology, and the correct development of immune cell populations. Several cytokines and growth factors utilize multiple intracellular signaling pathways such as the JAK-STAT pathway to transmit their information into the cell nucleus 18.
As mentioned, the SOCS proteins negatively regulate JAK-STAT signal transduction by forming a negative feedback loop that modulates cytokine-induced signaling through different mechanisms 19,20. The expression of SOCS3 gene is induced by phosphorylated STAT3 in response to cytokines that signal through gp130-related cytokine receptors, which in turn leads to attenuate or terminate the activation of STAT3 through targeting the receptor associated JAK kinases for degradation 9.
IL-6/STAT3 was an important pathways for the progress of IBD 21. Expression of SOCS3, which is a direct target gene of STAT3 and mediates suppression of IL-6 signaling through the ubiquitination and degradation of signaling intermediates 22. Mice with a deletion of SOCS3 in hematopoietic cells (Socs3fl/fl vav cre) have been shown to develop a severe inflammatory disease during adult life in response to pro-inflam-matory cytokines including IL-6 18. Accordingly, adenoviral-delivered SOCS3 reduced joint inflammation in mice with arthritis via inhibition of IL-6 signaling 18.
SOCS3 protein is an important regulator of cytokine signaling and consequently the immune response. Since SOCS3 regulates the function of a variety of cytokine associated with IBD, it plays a critical role in the development of these diseases 2. Studies in different mouse models have proven the critical importance of SOCS3 in restraining inflammation and allowing optimal levels of protective immune responses against infections 18.
SOCS3 promoter methylation is a major regulator of SOCS3 expression in the colonic epithelial compartment. Increased methylation in inflamed CD may result from the upregulation of DNMT1 expression. It thus seems that SOCS3 promoter methylation is an important mechanism controlling IL-6-dependent SOCS3 expression in colon cancer cultures 22. Data suggest that IL-6-dependent DNMT1 induction mediates SOCS3 downregulation, in turn releasing IL-6 signaling towards STAT3 from its negative regulation. A prediction from this hypothesis is that inhibition of DNMT1 should restore IL-6-dependent induction of SOCS3 and counteract methylation of SOCS3 promoter. Concomitant with the loss of SOCS3, STAT3 activation increased, providing a rational explanation as to why SOCS3 loss is related to an increased propensity to develop UC-CRC. Thus, we proposed that the loss of SOCS3 expression represents a critical event, permissive for the development of UC–CRC, but the molecular mechanism as to why chronic inflammation would provoke the loss of SOCS3 expression remained obscure at most 22.
Interestingly, when analyzing methylation status of SOCS3 gene, we observed very low levels of unmethylated DNA in both groups. The maximum inhibitory activity of SOCS1 and SOCS3 proteins toward the activation of STAT1 can be achieved by very low levels of SOCS protein expression, suggesting that SOCS proteins have a high affinity toward Jaks in vivo 23.
Conclusion :
In summary, this case-control study detected new profiles of CpG hypermethylation of SOCS3 gene that were associated with the susceptibility to CD development. Our study revealed a high negative correlation between promoter methylation and basal SOCS3 levels. We report that the hypermethylation status of SOCS3 in colorectal tissue specimens of CD patients is significantly higher compared to healthy controls. Our findings suggest that the CpG hypermethylation at promoter region of gene might influence SOCS3 gene expression and are fundamentally related to the etiology of UC. Our data provide an important insight into the STAT/SOCS axis by representing that epigenetic modifications in the SOCS3 gene regulation are the basis for hyperactivity of the inflammation-related IL6-STAT3 in UC patients. Further studies using in vivo and ex vivo systems will be needed to confirm our findings and even propose that how this knowledge may open novel avenues for the rational treatment of IBD.
Acknowledgement :
This study was supported by a grant from Tehran University of Medical Sciences (93-02-30-25932).
Ethics Approval :
The study protocol was approved by the Ethics Committees of Tehran University of Medicine science. Written informed consent was obtained from all individuals before sampling.
Conflict of Interest :
The authors declare that they have no conflict of interest.
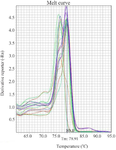
Figure 1. The figure shows the fluorescent melting peaks for the promoter CpG island of the SOCS3 gene. DNA melt curve analysis of changes in CpG methylation status was analyzed based on amplification of unmethylated bisulfite-treated DNA from colorectal biopsies of CD cases.
|
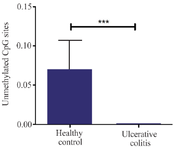
Figure 2. Comparison of changes in CpG island methylation status at the SOCS3 gene promoter region in colorectal tissue specimens of patients with ulcerative colitis and healthy control. Error bars mean± SD (*p<0.05).
|
|