Non-Invasive Detection of Esophageal Cancer using Genetic Changes in Circulating Cell-Free DNA
-
Ghorbian, Saeid
-
Department of Biology, Science and Research Branch, Islamic Azad University, Tehran, Iran
-
 M. Ardekani, Ali
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, Tel: 98 21 22432022 Fax: 98 21 22432021 E-mail: Ardekani@avicenna.ac.ir
M. Ardekani, Ali
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, Tel: 98 21 22432022 Fax: 98 21 22432021 E-mail: Ardekani@avicenna.ac.ir
-
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran
Abstract: Cell free DNA (cfDNA) is a genetic biomarker that is present in serum or plasma in high concentration in many types of cancer. Identification of circu-lating cancer related DNA molecules in serum or plasma is a non-invasive tool for early diagnosis and prognosis in many cancer patients. For this review, study selection and data extraction were performed by the authors. Detection of point mutations, microsatellite alterations, DNA hypermethylations and losses of heterozygosity in circulating cell free DNA have been characterized in esophagus cancer. Application of circulating cell free DNA as a biomarker, provide the best opportunity for constructing non-invasive tests for early de-tection, prognosis and management of cancer patients, after therapy in many types of cancer.
Introduction :
The esophagus is a muscular tube that con-nects the throat to the stomach. Cancer of the esophagus, also called esophageal cancer, can occur any place along the liner of the tube. Among the diverse types of existent neo-plasms, esophageal carcinoma is the eighth deadliest malignancy worldwide. Adenocar-cinoma and squamous cell are liable for more than 95 percent of all esophagus cancers (1).
The predominant histological subtype in esophageal cancer is squamous cell carcinoma (SCC) which contributes to about 80% of all esophageal cancers in the world (2). SCC has the highest incidence in Western countries with traditions of alcohol consumption, smok-ing or tobacco, hot drinks and malnutrition (3-5). ADC is found in industrialized countries, with gastric esophageal reflux (which causes Barrett´s Esophagus), obesity, substances de-rived from the grain of moldy corn (fumo- nisins), alcoholism, and smoking (Table 1) (3,6,7).
Epidemiology and Pathology of Esophageal Cancer :
Epidemiology
Epidemiologic data have shown variability in determining attitude in incidence of esoph-ageal carcinoma malignancies worldwide. The ascending incidence of esophageal cancer over the past two decades conformed to the change in histological type and primary tumor location. An estimated 14,500 deaths from esophageal cancer occurred in the United States in 2010 (8).
The American Cancer Society's most recent estimates for esophageal cancer in the United States for 2011 are as follows: about 16,980 new esophageal cancer cases diagnosed (13,450 in men and 3,530 in women) and about 14,710 deaths from esophageal cancer (11,910 in men and 2,800 in women). This disease is 3-4 folds higher among men than women (9). The incidence of esophageal can-cer varies remarkably with geographic region,
especially in developing nations. ADC is the most common type of esophageal malignancy in the United States and Western Europe (10). In America, the reported incidence of esoph-ageal cancer in patients is 3.2-4.0 per 100,000 persons (11), while in Europe and Asia, reports are 0.5-7.0 and 0.02-0.4 per 100,000 persons, respectively (12,13).
The incidence of SCC among the Asian countries is higher than that of ADC, espe-cially in countries and areas of East Asia such as Korea and Japan with incidence of 8.2-21 per 100,000 persons (14,15). Noticeable is the observation that the highest rates occurred in northern China and northern Iran, with inci-dence of about 40 per 100,000 persons in 2003 (13,16).
Pathology
Esophageal cancers are usually found with signs and symptoms that a person is having. The most common symptoms of esophageal cancer include: weight loss, difficulty in swal-lowing, chest pain, constant cough, bone pain, hiccups and pneumonia (17). These symptoms are usually present for several months before medical treatment is sought and initially pre-sents itself by having difficulty in swallowing dry foods. Furthermore, weight loss of 10 per-cent of normal body weight occurs in less than six months. About half of esophageal cancer patients present with locally advanced unresectable disease or distant metastasis.
ADC spreads via transverse penetration through the full thickness of the wall, whereas SCC tends to spread linearly in a submucosal fashion (18). Esophageal cancer spreads through extensive lymphatic channels with a skip me-tastases pattern when observed in autopsy specimens (19).
Staging in esophageal cancer
The diagnostic evaluations of ADC and SCC are essentially identical. Several strat-egies and approaches are acceptable for diag-nosis, staging, chemoradiotherapy, treatment, and surveillance of patients with esophageal cancer. These include endoscopy (20), bron-choscopy (21), thoracoscopy (22), laparoscopy (23), computed tomography (24), and surgery (25). However, maybe some of these approaches and strategies are associated with painful ex-periences and cause discontent for cancer patients.
Recently, research focus has been in the development of non-invasive screening at the early stages of cancer. In most cases, early detection of cancers is extremely difficult be-cause there are multiple uncertainties related to the location of the disease but it is neces-sary for full treatment and recovery. Current-ly, among the various approaches used for screening cancer patients, circulating molecu-lar biomarkers have been found to be of the most convenient and useful non-invasive tool with esophageal carcinoma (26).
Molecular genetic changes in esophageal cancer
The most common genetic alterations in esophageal cancer involve genetic alterations in several oncogenes, tumor suppressor genes, and DNA repair genes. Furthermore, the most common alterations found in esophageal can-cers such as deletions, amplifications and Loss Of Heterozygosity (LOH) has been re-ported to occur on several chromosomes list-ed in table 2 (27).
Mutations in oncogenes and tumor suppressor genes
Tumor suppressor genes p53 and p16 have been introduced to the primordial G1 cell cycle regulatory genes (28). Alteration of these genes such as LOH and mutation can lead to the loss of regulation of cell growth, which is important in carcinogenesis (29). Mutation of p53 has been extensively investigated in both SCC and ADC (30,31). Previous studies have reported the incidence of p53 mutations (53 percent in ADC and more than 93 percent in SCC) (32) especially, in the exons 5-8 (encod-ing the DNA-binding domain of the protein) in SCC. In addition to mutations in p53, LOH at chromosome 17q13 is a significant event in both SCC and ADC.
In principle, the hypotheses were that p53 alteration occurs frequently as an early event in the tumor progression of esophagus carcin-oma (33). However, inactivation of the p16 gene, LOH, genetic mutation and aberrant DNA methylation in the coding and non-coding (promoter) regions also frequently occur in both SSC and ADC (34,35). Tarmin et al, has reported the incidence of LOH in about 50 to 65 percent of patients with SCC at the 9p21 chromosomal locus (36). However, aber-rant p16 hypermethylation in the promoter region is a common mechanism for the inacti-vation of this gene in SCC. Taghavi et al, showed the aberrant hypermethylation of this gene in 62 percent of SCC patients in Iran (37).
Previously, Abbaszadegan et al, had report-ed incidence of this aberration in SCC to be 73.3 percent in the northeastern Iran (38), while Hardie et al, reported that hypermethylation of the p16 promoter is 85 percent in ADC (39). Furthermore, inactivation of p14 and p15 genes has been observed in esophagus cancer. Aberrant methylation in the promoter regions of these genes was reported to be associated with loss of transcription in previous studies (40,41). Xing et al, reported alteration of methyl-ation patterns in the promoter of p14 and p15 genes and incidence of LOH in p14 and p15 genes in SCC tumor samples (42).
The Adenomatous Polyposis Coli (APC) gene is a tumor suppressor gene located at chromosome 5q21 and is involved in esoph-agus cancer (43). One of the alterations in this gene is LOH at 5q, occurring in about 55-80 percent of SCC and 20-55 percent of ADC (44,45) cases, but mutation of APC is under 10 percent in this cancer type (46). The most com-mon type of gene inactivation occurs via hypermethylation in the promoter region of the APC gene with an incidence of 92 percent in ADC and 50 percent in patients with SCC (47). Therefore, the hypermethylation of this gene has noticeable roles in both SCC and ADC.
The cyclin D1 gene, located at chromosome 11q13, encodes a protein that is required for controlling the cell cycle (48). Previous studies have analyzed cyclin D1 expression in patients with esophagus cancer and aberrant overex-pression of this gene in 23-73 percent of pa-tients with SCC has been reported (49,50). Furthermore, Metzger et al, showed an overe-xpression and gene amplification of cyclin D1 in 22-64 percent of patients with ADC (27).
The erbB-2 (HER2) is one of the members of Epidermal Growth Factor Receptor (EGFR) family, which acts as tyrosine kinase receptor. This receptor has an intracellular tyrosine ki-nase activity and extracellular binding do-mains. The erbB-2 oncogene encodes a trun-cated form of EGFR which contains continu-ous tyrosine kinase activity (51). The overex-pression and gene amplifications of erbB-2 has been demonstrated in patients with SCC and ADC with 20 to 60 percent frequency (52).
Microsatellite instability
Microsatellites are short tandem repeats of nucleotide sequences found at about 5000 base pair intervals. Microsatellite Instability (MSI) has been recognized as a length muta-tion that occurs especially in microsatellites. The two genes studied in this alteration in-clude MSH2 and MLH1 which are located at chromosomes 2p and 3p, respectively. The functions of these genes are essential in DNA mismatch repair and reduction of genomic replicative error rate. Several studies have reported 10 to 20 percent frequency of MSI in patients with ADC and SCC, with a higher frequency in SCC patients (53-55).
Molecular Biomarkers :
Circulating molecular biomarkers
Currently, progress of proteomics has open-ed the door to cancer-related biomarker dis-covery. Proteomics is the complete descript-tion of all proteins encoded by the genome, called the proteome (56). Advances in proteo-mic technologies such as the development of quantitative proteomic methods, high-reso-lution and high-throughput methods have been used to identify and understand patho-physiology of carcinomas. Rapid advances in the field of proteomics promise discovery of biomarkers which could potentially aid in early diagnosis, prognosis and accurate pre-diction of outcomes during cancer treatments and management (57-62).
Fortunately, biological fluids such as urine, blood, serum or plasma contain many bio-markers originating in many different tissues of the body. In the past few decades different biomarkers have been tested as screening tools for cancer patients. Proteins are prob-ably the first-generation of biomarkers to have been investigated and discovered in the biological fluids. For example Prostate Spe-cific Antigen (PSA), CA19-9, CA125, squa-mous cell carcinoma antigen (SCC) and cyto-keratin 19 fragment (CYFRA) are clinically applied as biomarkers (63-66). However, these conventional biomarkers have low sensitivity and insufficient specificity. Thus, researchers have looked toward the other types of tumor biomarkers (67).
Recent advances in analytical assay tech-nologies have allowed development of ampli-fication techniques that are based on circu-lating nucleic acids (RNA and DNA) as bio-markers in serum or plasma of cancer patients (68). Over the past decade, tremendous amount of information has been accumulated which are related to cancer-specific changes such as gene mutation, Single Nucleotide Polymorph-ism (SNP), gene deletion, LOH, epigenetic al-terations (69), genome instability (70) and aber-rant at the expression level at RNAs and pro-tein levels (71,72).
In recent years, efforts in many laboratories throughout the world have been focused on the utilization of genetic and expression ab-normalities in circulating biomarkers for early detection, prognosis and assessment of cancer patients after therapy (73).
Biological characteristics of cfDNA
In 1948, Mandel and Metais reported the existence of circulating extracellular nucleic acids in human blood (74). Thereafter, Stroun et al, showed biological characteristics of circulating cell free DNA (75). Circulating cfDNAs are small fragments of genomic DNA present in the plasma or serum (76). The mechanisms of the occurrence of cfDNA in blood under normal and pathological condi-tions remain unknown. Two main mechan-isms of release cfDNA in the plasma or serum have been postulated: (a) cells detach and extravasate into the blood stream where they undergo lysis and (b) cells undergo apoptosis or necrosis in cancer tissues in vivo and their DNA is released in the blood stream (77). Both molds simultaneously contribute to cfDNA production in cancer patients.
In the programmed enzymatic apoptosis, low molecular weight DNA fragments of about 185-200 base pairs are released in the necrosis products, much of the high molecular weight DNA fragments of about 400 base pairs were detected in serum and plasma (78,79). The role of macrophages in degradation of DNA fragments and then release into the bloodstream has been suggested to probably occur through engulfing process of necrotic or apoptotic cells (80).
Isolation and quantification of cfDNA
The cfDNA is extracted using phenol/ chloroform or standard commercially avail-able DNA kits such as glass-milk-based methods, nucleospin blood kit and PureGene DNA isolation kit from serum or plasma tumor patients. Furthermore, a variety of DNA quantification methods is utilized to measure including DipStick (Invitrogen), fluorometry with SYBR-green and real-time PCR (76). Diehl et al, developed a technique called BEAMing (beads, emulsion, amplification and magnetics) for quantification of circu-lating mutant cfDNA in serum or plasma of tumor patients (81). However, with the appear-ance of novel approaches incorporating auto-mated DNA isolation and amplification with multiplex quantitative fluorescent PCR of circulating cfDNA, the potential to utilize cfDNA as a screening tool for many types of cancer exists.
Non-Invasive Screening of Esophagus Cancer using cfDNA :
Alterations at the cfDNA level
Leon et al reported that the average cfDNA concentration in the serum or plasma of can-cer patients is higher than in normal subjects (82). This is because the increase of cfDNA levels in the serum or plasma of tumor pa-tients are contributed by both tumor DNA and non-tumor DNA. However, meta-analysis data has shown high discrepancy in the cfDNA concentrations between studies. And this probably depends on variables such as number of patients in a given study, cancer subtypes, tumor size, stages, location, and other risk-related factors (75).
Unfortunately, high variability in the abnor-mal cfDNA concentrations has prevented it from becoming more than a critical biomarker in each of malignancies (83). One of the com-plicating facts is that an increased plasma cfDNA level has been also observed due to inflammation (84), trauma (85), premalignant states (86), after exercise (87), and in patients suffering from acute or chronic illnesses (88,89).
Several investigators have reported circu-lating cfDNA levels to be significantly higher in patients with malignancies such as lung, breast, colon, hepatocellular, ovarian, pro-state, and melanoma than in healthy individ-uals (90-93). In another study, Banki et al, re-ported that cfDNA level in patients with esophageal cancer was significantly higher and after complete resection of the tumor, the cfDNA level returned to normal (94). Chikawa et al, reported similar results in which the average circulating cfDNA concentrations from esophageal cancer patients were shown to be significantly higher than those in healthy controls (95). An earlier study had shown that the average concentration of cfDNA in serum was 13 ng/mL in a healthy individual and
180 ng/mL in various cancer patients (80). De-tection of differences in cfDNA levels may be a good approach and potentially a valuable tool for early detection and for the evaluation of the prognosis of patients with esophagus carcinoma.
Methylation analysis
Aberrant DNA methylation patterns as a measurement of epigenetic changes are fre-quently found in several types of cancer. The epigenetic changes are described as changes in hypomethylation and hypermethylation levels in special gene regions (e.g. promoter regions). For example, tumor suppressor genes have been demonstrated to be hyper-methylated at an early stage of tumorigenesis and hypermethylation process is known to silence regulatory genes involved in the cel-lular pathways related to cancer. Esteller et al, proposed detection of aberrant methylation changes of cfDNA in the serum or plasma in tumor patients and suggested it can be used as a tool for early detection and monitoring of the efficacy of therapy in tumor patients (96).
Kawakam et al, observed hypermethylation in APC DNA, with incidence of 6.3 percent in the plasma of SCC patients and 25 percent in plasma of ADC patients (97). Furthermore, Hibi et al, detected aberration in methylation level of the promoter of p16 gene in serum DNA of 18 percent of SCC cancer patients (98). Liu et al, assessed the methylation status of Wnt antagonist family of genes in esoph-ageal cancer patients and found that hyper-methylation of promoters for SFRP-1, WIF-1, DKK-3, and RUNX-3 genes could be detected in plasma DNA using Methylation-Specific PCR method (99). Therefore, detection of aber-rant promoter hypermethylation of cancer re-lated genes in serum may be useful for esoph-ageal cancer early diagnosis and detection of recurrence.
Microsatellite analysis
The presence of microsatellite alterations in the serum or plasma could be used as a prog-nostic indicator. Microsatellite analysis of cir-culating cfDNA represents an appearing group of biomolecular tumor markers, where as control subjects show no serum alterations (100). Claus et al, analyzed several micro-satellite markers which are commonly altered in serum of patients with SCC of the esoph-agus. These markers were located at chromo-somes 9p (p16), 17p (p53) and 5q (APC gene) and he observed that in 96.4 percent of cases, at least one alteration was found in the serum DNA specimen of esophagus cancer patients (101). Identification and analysis of microsatel-lite alterations in the serum DNA from SCC patients may be a valuable tool for evaluation at early stage and follow-up studies.
Conclusion :
Early diagnosis of cancer provides much promise for full recovery of patients. Unfortu-nately, conventional methods of cancer scre-ening are often invasive and expensive. Sensi-tivity and specificity of these methods are also insufficient for diagnosis of cancer at an earlier stage. For this reason, many research-ers are attempting to increase sensitivity and specificity of methods for early detection and monitoring of tumor recurrence.
In the past decade, there has been a revo-lution in the number of studies analyzing cfDNA of tumor, as a reliable substitute for tissue analysis and a possible tool for the prognosis and diagnosis of cancer patients. At the present time, the major advantage of cfDNA as a biomarker, is the easy accessibil-ity and its stability in plasma or serum speci-mens.
Another advantage is the use of minimally invasive method for obtaining blood to ana-lyze cfDNA and this may have the potential to replace the existing cancer tissue biomark-ers in the future. The combined observations that a correlation exists between molecular al-terations such as mutations, microsatellite in-stability and aberrant methylations in cfDNA and clinical data in esophageal cancer pa-tients, is a strong indicator that cfDNA would likely play a role in early diagnosis and prog-nosis of esophageal cancer patients in the future.
Furthermore, recent developments of se-quencing methods and characterization of cir-culating cfDNA obtained from a variety of cancer patients in different conditions would increase our knowledge about the mechan-isms and functions of circulating cfDNA. Ex-ploration for novel cancer biomarkers with improved diagnostic sensitivity and specifi-city should help clinicians to apply therapeut-ics more efficiently and effectively during the management of the cancer disease.
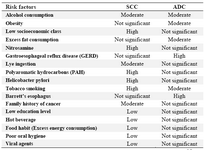
Table 1. The most common risk factors affecting the development of esophageal carcinomas
SCC, Squamous cell carcinoma; ADC, Adenocarcinoma. Information from references (3-7)
|
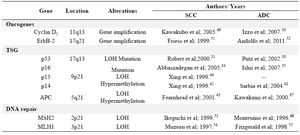
Table 2. Summary of the most common alterations in esophageal cancers
SCC, Squamous cell carcinoma; ADC, Adenocarcinoma; LOH, loss of heterozygosity; TSG, tumor suppressor gene; APC, adenomatous polyposis coli; MLH1, mutL homolog1
|
|