Factor VIII as a Novel Biomarker for Diagnosis, Prognosis, and Therapy Prediction in Human Cancer and Other Disorders
-
Khalilian , Sheyda
-
Student Research Committee, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
Department of Medical Genetics, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
USERN Office, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
Mohajer , Zahra
-
Student Research Committee, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
USERN Office, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
 Ghafouri-Fard, Soudeh
Department of Medical Genetics, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran, Tel/Fax: +98 21 23872572; E-mail: s.ghafourifard@sbmu.ac.ir
Ghafouri-Fard, Soudeh
Department of Medical Genetics, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran, Tel/Fax: +98 21 23872572; E-mail: s.ghafourifard@sbmu.ac.ir
Abstract: Coagulation factor VIII (FVIII) is an essential cofactor in the coagulation cascade, encoded by the F8 gene on the long arm of chromosome X (Xq28). FVIII is normally circulated in complex with Von Willebrand factor (VWF) and has relevant emerging extracoagulative functions. Dysregulation of FVIII is associated with tumor progression, and could be used as a novel biomarker for tumor screening and monitoring. In breast cancer, bladder cancer, colorectal carcinoma, esophageal carcinoma, hepatocellular carcinoma and lung cancer, F8 is regarded as an oncogene. In coronary heart disease, hemophilia A and liver disease, F8 dysregulation has been recognized as a potential biomarker for disease diagnosis and prognosis. However, the basis of these differential expression levels remains to be understood. In this review, which is a mixture of literature review and bioinformatics analysis we described the biological functions and characteristics of FVIII, and also its expression level in non-malignant disorders and various cancers.
Introduction :
Coagulation factor VIII (FVIII), a significant multi-domain glycoprotein component of the intrinsic coagulation pathway, consists of six domains: heavy chain (A1-A2) and light chain (A3-C1-C2) 1. The coagulation FVIII is encoded by the F8 gene on the long arm of chromosome X (Xq28). F8 gene consists of 26 exons and is approximately 186 Kb 2,3. The 3D structure of FVIII is constructed by the SWISS PDB viewer (https://www.rcsb.org/3d-view/3CDZ/1), and is shown in figure 1A. There are several proteins that interact with FVIII. The protein-protein interaction of FVIII is analyzed by STRING (https://string-db.org), and is represented in figure 1B. Mounting evidence states that there are over 3000 mutations in F8 gene, including small deletions, insertions, large deletions, nonsense, and mis-sense (substitutions). Additionally, intron 1 and 22 inversions make up approximately 50% of total mutations 4,5.
FVIII initially binds to von Willebrand factor (vWf), which is a serine protease in plasma that preserves FVIII from degradation and clearance by Dendritic Cells (DCs). In response to injury, FVIII is cleaved by thrombin and forms activated FVIII (FVIIIa) which is then separated from vWf and joins the activated platelets via C1-C2 domains. FVIIIa binds to activated FIX (FIXa), another coagulation factor to generate intrinsic tenase complex. This complex then leads to the formation of activated factor X (fXa) and also thrombin. Free FVIII is an unstable molecule with a labile structure with a half-life of about two hours. Generation of the FVIII-vWF improves the FVIII half-life time in the blood circulation 6. FVIII and VWF production are not localized in the same tissues. While sites of VWF production have been discovered across the vascular endothelium, especially in the lung and brain, FVIII is largely generated in the liver, specifically in sinusoidal endothelial cells with almost different expressions 7-10. In addition, the expression of FVIII is different in various tissues. Overall, the biological functions, and also expression changes of FVIII remain unknown during decades.
Several investigations have reported high levels of circulating FVIII in cancer patients 11-13. Notably, FVIII is independently associated with increased risk of venous thromboembolism in these patients. In evaluating the potential causes of increased circulating FVIII levels in cancer patients, the following parameters should be considered: the physiological and cancer-related sites of FVIII production, and the body’s response to cancer. The latter has several shared components with wound healing and inflammatory proteins 11-13.
In this review, we have mainly focused on the differential expression of FVIII in human disorders, especially cancer, to improve our understanding of its biological functions, and importance in disorders. Also, the FVIII potential to be identified as a novel diagnostic, prognostic, and therapeutic biomarker has been clarified.
The Role of FVIII in Human Diseases (Table 1) :
Non-Malignant Diseases
Coronary Heart Disease: In multiple prospective researches, the function of FVIII as a risk factor for incident Coronary Heart Disease (CHD) and stroke has been hypothesized. However, it is still unknown if the FVIII connection is independent of the other Cardiovascular Disease (CVD) risk factors 14-18. FVIII is significantly linked to risk factors associated with atherosclerosis, including age, high Body Mass Index (BMI), diabetes, and inflammatory mediators 19.
In an experiment performed by Raffield et al 20, 3493 African-American patients were studied in the Jackson Heart Study. According to this study, compared to other subclinical outcomes, elevated FVIII was substantially linked to Left Ventricular Hypertrophy (LVH) (p=0.01). In Cox models that were moderately modified for gender and age, sustained FVIII level was substantially linked with general hard CHD, Heart Failure (HF), and death, in contrast to stroke. After adjusting for conventional CVD risk factors, such as CRP, these relationships were thus diminished. Raffield et al observed a model adjusted for CVD risk factors, such as CRP. The results showed that the p values for the correlation with HF and death were significant (p<0.05); while the correlation with hard CHD was marginally significant (p=0.05). Numerous other multiracial investigations have produced results indicating that FVIII is more strongly correlated with mortality than incident CVD 14,21.
Hemophilia A: Hemophilia A (HA) is an inherited X chromosome-linked hemorrhagic condition that either results in the down expression of the FVIII gene or the synthesis of an abnormal FVIII protein 22,23. Jankowska et al 24 studied the HEK-293T cell line using an MS2-TRAP assay. In order to conduct this experiment, they transfected HEK-293T cells with either MS2 alone (negative control) or MS2 fused to the FVIII. The RNA samples were isolated from cells transfected with MS2 alone and MS2 fused with FVIII. The second batch of cells had a 5-fold increase in miR readings overall. Of the 64 miRNAs that Jankowska et al 25 found to be linked to the FVIII mRNA, 22 likewise expressed at higher levels (higher than 2-fold) in HA patients compared to healthy controls, and 8 were observed at lower levels (less than 0.5-fold) in HA patients, particularly miRNA-19b-3p and miR-186-5p. These two miRNAs were also detected in the experimental pull-down assay with the highest read count. One explanation for high FVIII levels is the downregulation of the miRNAs that typically suppress the FVIII gene. Moreover, upregulation of the so-called miRNAs can reduce the expression of FVIII, leading to severe HA 26-29.
In another study by Sarachana et al 30, 15 HA patients were examined. They extracted the total RNA from the whole blood sample collected from HA patients and the healthy control group. Then they carried out an analysis of non-coding RNA (ncRNA) expression profiling, observing that numerous ncRNAs are dysregulated in HA patients. The hsa-miR-1246 was found to be the most significantly dysregulated miR. Sarachana et al also found a hsa-miR-1246 binding site on the FVIII gene, suggesting that it can regulate the FVIII gene expression. Their experiment showed that overexpression of has-miR-1246 leads to suppression of FVIII, although the effect of this ncRNA on the clinical manifestations of HA varies in different patients. Moreover, some of the other overexpressed miRNAs seen in HA patients include hsa-miR-374b-3p, hsa-miR-5581-3p, hsa-miR-6803-3p, hsa-miR-30c-3p, and hsa-miR-542-3p. Mutations in the 3′UTR of FVIII alter the process of splicing and expression of mRNAs, resulting in downregulation of FVIII, which can cause HA 31.
Additionally, Mei et al 32 experimented on several cell lines to identify the most efficient for the FVIII gene expression. The results showed that the FVIII levels in the HKB11 cells were significantly higher than the other cell lines, both intracellular and on the cell surface. Mei et al also investigated the HKB11 cells to discover the mechanism of which the FVIII expression is regulated. HKB11 cells were a hybrid of human embryonic renal cells and human B lymphocytes, suggesting that the overexpression of FVIII gene in these cells is due to HKB11 cell inheriting the embryonic characteristics of kidney in transcription and translation of FVIII mRNA, and also the characteristics of B cells in the process of protein secretion.
Liver disease: Liver disorders are linked to significantly high levels of FVIII, while several other proteins and coagulative factors are decreased. Although, the pathological process of FVIII elevation is still unclear 33,34. In a study by Hollestelle et al 33, 19 patients with liver disease were studied. The FVIII level was increased in 13 out of 19 patients. Also, individuals with hepatic cirrhosis had significantly lower levels of FVIII mRNA than control patients (p=0.01). Therefore, it would seem that rising levels of FVIII did not correspond to rising FVIII gene transcription; instead, it appears that there was an inverse correlation observed, but it was not significant. Additionally, there were no variations in the cellular distribution pattern of FVIII between patients. Nonetheless, they observed that bigger vessels in cirrhotic tissue appeared to overgrow sinusoidal endothelial cells that generate FVIII. This could clarify why FVIII mRNA expression was lower in the cirrhotic tissues than in non-cirrhotic. Raffield et al also evaluated at the expression of VWF at the mRNA and protein levels to gain understanding into the seemingly different cellular expression of FVIII gene and plasma FVIII levels. It is widely known that VWF plays a key role in controlling plasma FVIII concentrations. FVIII is protected by VWF from premature clearance and proteolytic degradation. Therefore, according to the increase in VWF in liver disease, it is obvious that overexpression of VWF leads to an elevation in FVIII levels 35-37.
Cancers
Breast cancer: Several previous studies have highlighted the prognostic effects of coagulative factors in Breast Cancer (BC) treatment outcomes. Mandoj et al 38 conducted a study on 235 patients with BC staged I to IIA. The serum level of coagulation activation factors including FVIII were measured before therapeutic procedures. The levels of FVIII in patients with early stages of BC were considerably elevated. Mandoj et al found that high levels of FVIII are correlated with intermediate mortality risk 38.
Bladder cancer: It has been demonstrated that cancer-related thrombosis with high risks of Venous Thromboembolism (VTE) happens in Bladder Cancer (BLC) 39,40. Although the mechanism is still somewhat unclear, FVIII is believed to be an independent risk factor for VTE in several cancers 20-22. Walker et al 41 performed an experiment on the cell line ECV-340. They found that in 7 out of 9 urothelial carcinomas the FVIII levels were elevated in the cancer cells. The highest expression of the FVIII gene was detected in invasive BLC cells. Also, the FVIII gene was found to be overexpressed in the cancer cells compared to the healthy adjacent tissue. Walker et al chose the BLC cell lines for the experiment because of the high VTE incidence rate in the BLC cells 39,40.
Triple-negative breast cancer: Gujam et al 42 studied 360 patients with invasive ductal BC to determine the prognostic value of Lymphovascular Invasion (LVI) and Blood Vessel Invasion (BVI) in node-negative and triple-negative BC. The experiment showed that both lymphatic and blood vessels were persistently positive for FVIII. The results suggested FVIII as a predictive factor for tumor recurrence, cancer-specific survival rate, and metastatic node formation.
Colorectal carcinoma: Schellerer et al 43 studied 79 patients with Colorectal Cancer (CRC). Following measuring the plasma levels of FVIII, they found an elevation in the FVIII level in CRC cells compared to healthy cells. Schellerer et al detected a significant elevation in the FVIII levels in the cells with stage II-IV cancer, while cells with stage I cancer showed FVIII levels in the normal range. Their results showed that staging and differentiation of the tumor have significant impacts on the FVIII levels. But there was no connection between FVIII levels and the occurrence of metastases.
Esophageal carcinoma: Byrne et al 44 performed a study on 50 patients with Esophageal Cancer (EC) after the chemoradiotherapy cycles. Measurement of FVIII levels showed a significant elevation in FVIII levels following the chemoradiotherapy sessions. Additionally, increased FVIII levels might cause primary or recurrent VTE. Although the exact pathogenetic process of this elevation is still unclear.
Hepatocellular carcinoma: Zhuang et al 45 conducted an experiment on the human cell lines Hepal-6, HepG2, and HUVEC, and the mice cell line C57BL/6J to determine whether the dydio controlling coagulative biomarkers plays a role in the metastasis of Hepatocellular Carcinoma Cells (HCC). Dihydrodiosgenin, a parent aglycone of diosgenyl saponin, commonly known as dydio, has anticancer, anti-inflammatory, and antithrombotic mechanisms 46-48. Diosgenin derivatives have been shown in prior investigations to considerably lower FVIII levels in animal models 49,50. Moreover, liver diseases such as HCC are correlated with high levels of FVIII 33. Accordingly, it appears that dydio could control the progress of HCC by regulating the level of FVIII. Zhuang et al demonstrated that dydio decreased HCC metastasis by preventing platelet activation and lowering FVIII levels 45.
Lung cancer: In a survey by Liu et al 51, 115 patients with Lung Carcinoma (LC) were studied. Blood samples were collected for measuring the plasma levels of coagulative factors. The results showed a significant elevation in FVIII levels in LC patients compared to healthy individuals regardless of existent metastasis. These findings are in accordance with prior research that claimed patients with metastatic diseases had a higher propensity to develop coagulative and fibrinolytic disorders 52-54. Even though the biological relevance of hemostatic disorders in cancer is yet unknown, findings demonstrate that activation of the coagulative-fibrinolytic pathway by cancer cells may enhance invasiveness and metastases 55. Additionally, higher coagulative biomarker levels have been linked to a poor prognosis in LC and failure to treatment 56. Table 1 represents the expression changes and roles of FVIII in human non-malignant and malignant disorders.
Database analyses
GEPIA2: GEPIA2 (Gene Expression Profiling Interactive Analysis 2) database, analyzes the data of RNA sequencing expression for approximately nine thousand normal and tumor samples from the projects GTEx and TCGA with the help of standard processing pipelines 57. We examined the connection between the overall prognosis and the F8 expression levels in a variety of cancer patients. We also used this database to investigate the link between the expression of FVIII and immunological responses and tumor markers in particular. Using GEPIA2 databases, the expression of F8 was significantly increased in normal controls than in tumor tissues in Bladder Urothelial Carcinoma (BLCA), Breast invasive Carcinoma (BRCA), Colon Adenocarcinoma (COAD), Kidney Chromophobe (KICH), Lung Squamous Cell Carcinoma (LUSC), Ovarian serous cystadenocarcinoma (OV), Rectum Adenocarcinoma (READ), Testicular Germ Cell Tumors (TGCT), Uterine Corpus Endometrial Carcinoma (UCEC), and Uterine Carcinosarcoma (UCS). In contrast, the expression of F8 was significantly decreased in normal control than in tumor tissues in Thymoma (THYM). Figure 2A represents the expression of F8 in different cancers based on GEPIA2 database.
In addition, GEPIA2 is a web tool for investigating the impact of genes on survival in different cancer types. A log-rank p-value <0.05 is considered statistically significant. As shown in figure 3, the survival analysis by the GEPIA2 database indicated that overexpression or down- regulation of FVIII was not associated with overall survival in lung, breast or BLCs.
TIMER: The TIMER database 58 was also used to compare the F8 expression in normal and tumor tissues, and the results are represented in figure 2B. Besides, we investigated the infiltration of immune cells in several tumors with the help of TIMER database. We included the data on immune infiltration in cancers, which were derived by statistical techniques and verified by pathological tests. We investigated the link between the expression of FVIII and the intensity of infiltration using particular immune cell subsets and various immune invasion assessment algorithms. Lastly, correlations between the expression of FVIII and the immune cells, including CD8+ T cells, macrophages, neutrophils, and Natural Killer (NK) cells were examined, and figure 4 represents the results. For instance, in BRCA, F8 had no significant correlation with macrophages and CD8+ T cells and a positive correlation with neutrophils, and NK cells, based on the MCP-counter algorithm.
cBioPortal: The cBioPortal database provides numerous options for exhibiting both discrete genetic events such as mutations and continuous events, including information on the quantity of mRNA or proteins or DNA methylation. The cBioPortal offers access to summarized data on every cancer study featured in the portal, along with performing individual gene searching 59. We assessed the F8 gene alterations using the data from the cBio Cancer Genomics Portal (http://cbioportal.org) database. As shown in figure 2D, the mutations and substitutions in the F8 gene were responsible for most changes in most cancers.
COSMIC: The COSMIC 60 (Catalogue of Somatic Mutations in Cancer) database stores somatic mutation information and related metadata. It is accessible through a series of online websites and offers multiple export choices as well as graphical or tabular views of the data. Each gene is kept in COSMIC in a static form. Ensembl 61 helped to determine the genomic structure of every gene and its chromosomal placement. The RefSeq project was used to determine the cDNA and protein sequencing 62,63. The COSMIC database was used to analyze the F8 mutation distribution in various tissues (Figure 5), which demonstrated that the F8 mutation rate was different in most tissues and disorders. Pie chart represents that mis-sense substitutions account for the largest amount of mutation types among total samples (46.58%) (Figure 5A). Additionally, the C>T mutation was the most common substitution mutation in the F8 gene (Figure 5B).
TargetScan and miRDB: TargetScan (https://www.targetscan.org/), and miRDB databases (http://www.mirdb.org/), are two powerful miRNA prediction tools, that were used to predict the miRNAs that target F8 gene. A total of 76 miRNAs have been identified that target F8 gene via these two databases (Figure 6). Based on the results, F8 is predicted to be targeted by a variety of miRNAs that are involved in a wide range of cellular functions.
Enrichr: Gene ontology enrichment analysis for the protein interaction network of FVIII was performed using the Enrichr database 64. Gene ontology is categorized into three separate groups: Biological Process (BP), Molecular Function (MF), and Cellular Component (CC). Table 2 shows the results of the GO annotation enrichment analysis related to the interaction network of the FVIII gene. The interaction network of the FVIII was remarkably enriched in GO terms, comprising the peptidyl-asparagine modification, regulation of blood coagulation, protein N-linked glycosylation via asparagine, zymogen activation, etc. in BP; serine-type endopeptidase inhibitor activity, endopeptidase inhibitor activity, protease binding, mannose binding, peptidase inhibitor activity, etc. in MF; COPII-coated ER to Golgi transport vesicle, intrinsic component of the external side of the plasma membrane, platelet alpha granule lumen, clathrin-coated vesicle, platelet alpha granule, etc. in CC. Additionally, the KEGG pathway analysis uncovered that platelet activation, Coronavirus disease, and ECM-receptor interaction pathways were the most significant pathways for the FVIII gene and its interaction network (Table 3).
Discussion :
Approximately 50% of cancer patients and more than 90% of those with metastatic illnesses show hemostatic disorders 65. Even though elevated VWF and FVIII levels have been detected in cancer patients, the precise mechanism relating the rise in the levels to VTE or survival rates is still up for debate 66. FVIII normally circulates coupled with VWF and functions as an acute phase reactant protein and an activator in the coagulation cascade. VWF prevents FVIII from proteolytic attacks and premature removal by binding to it 35-37,67-70. Typically changes in the plasma level of VWF will affect FVIII levels with the same change 71,72. Increased FVIII levels in cirrhotic patients are most likely caused by increased hepatic VWF production rather than enhanced liver FVIII overexpression activity. Additionally, upregulation of the FVIII gene can lead to recurrent VTE, and downregulation of the gene leads to X-linked hemorrhagic disorder HA 19,70,73,74. FVIII is significantly associated with risk factors of atherosclerosis, including age, diabetes, high BMI, and several coagulative markers 19. Increased levels of FVIII have been suggested as an independent risk factor for death overall as well as arterial thrombotic diseases including Myocardial Infarction (MI) and stroke 15-18,21,75,76. There is a much higher elevation in the FVIII levels in African-Americans (AA) than in patients of European ancestry (EA). Therefore, AAs may be at greater risk of VTE than EAs 77-80.
According to analyses in Jackson Heart Study (JHS), FVIII is significantly associated with age, diabetes, the CRP biomarker, and triglycerides in AAs. Moreover, FVIII is more significantly associated with mortality and less with CVD incident. The reason remains unknown. High levels of FVIII can be due to the downregulation of the miRNAs that typically bind to the FVIII gene and suppress it. On the other hand, overexpression of the so-called miRNAs can result in HA phenotypes. Our results have represented a total of 76 miRNAs that could target F8 gene and might be associated with pathogenesis of several disorders.
There are numerous ways to find and study the miRNA-mediated regulation of the gene expression. Samples from patients and healthy individuals were collected, and following the deep-sequencing process, the expression levels of miRNAs were observed. The MS2-TRAP assay helped with detecting the interactions between FVIII gene and miRNAs 26,29,81-92. The FVIII gene can be suppressed by both miR-186-5p and miR-19b-3p. Additionally, it was shown that a severe HA patient had considerably greater levels of miR-19b-3p expression 30. Furthermore, levels of FVIII mRNA expression in cirrhotic patients were lower than those of healthy individuals. There were no variations in the intracellular distribution of FVIII in the participants. Nevertheless, the bigger arteries in the cirrhotic tissue appeared to overgrow sinusoidal cells that generate FVIII. This could be the reason why FVIII expression levels were lower cirrhotic tissues than in healthy counterparts. Also, the amount of data for the link between coagulation abnormalities and cancer has substantially increased in recent years 93,94. Several coagulative biomarkers have been suggested to have prognostic values in many cancers independently, in the case of VTE 95-98. In women with recently diagnosed invasive BC, the plasma levels of FVIII strongly correlated with lymph node metastasis and the number of them, and also, with the HER2 status 99. Moreover, FVIII is correlated with cancer-specific mortality 21.
Furthermore, high VTE occurrence rates have been seen in BLCs, which is associated with the increase in the early biomarkers of coagulation 100,101. FVIII expression was seen to be notably higher than VWF and other coagulative markers. The invasive BLC showed the highest levels of FVIII expression when compared to other bladder tumors 41.
Studies have demonstrated that in colorectal cancers, the tumor's staging and differentiation have an impact on the plasma level of FVIII. FVIII levels, however, are not linked to the formation of metastases 43. It is noteworthy that three years following surgery, patients with lower FVIII levels at the time of the procedure had greater survival chances. In patients with EC after preoperative chemoradiotherapy, FVIII plasma levels are significantly raised. There was no discernible change in FVIII levels between the multimodal group and the patients who only underwent surgery 44. As mentioned above, VTE is commonly discovered in cancer patients, which adds to the morbidity of the patients. In around 50-80% of HCC patients a portal or hepatic vein invasion is detected, and tumor thrombus is known to indicate a bad prognosis for these individuals. When the tumor spreads and develops a tumor thrombus, it may be fatal 102-106.
According to current studies, patients with lung cancer had shown considerably higher plasma levels of many coagulative factors, including VWF and FVIII regardless of the presence of distant metastases. Additionally, increased levels of coagulative biomarkers have been associated with poor prognosis for lung cancer 51. Most importantly, in an attempt to assess association between FVIII and cancer, Walker et al, conducted a research to investigate the possible direct expression and secretion of FVIII by cancer cells. The goal of their research was to assess the expression and synthesis of FVIII in cancer, using BLC as the model system. They demonstrated that FVIII expression is elevated in BLC compared with normal bladder tissue, and it is released by BLC cells. Moreover, they showed that this can be extended to other cancer cell lines with a pattern independent of VWF and different relevant players in the coagulation cascade, thus providing criteria of a potential independent function for FVIII in cancer-related pathophysiology 11.
Conclusion :
In the present study, we have analyzed the expression of F8 in various tumors and adjacent normal tissues using publicly available expression databases. F8 was differentially expressed between normal and tumor tissues in different cancers. Furthermore, F8 expression levels were significantly related to levels of immune cell infiltration. These results suggest that F8 has a significant function in the regulation of immune cell infiltration in several tumors, with particularly high impact on macrophages, neutrophils, CD8+T cells, and NK cells infiltration. Therefore, these results indicate that F8 contributes to the immune response in various tumors and could be a novel prognostic biomarker.
Acknowledgement :
Not applicable.
Funding: Not applicable.
Conflict of Interest :
The authors declare they have no competing interests.
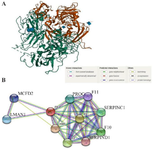
Figure 1. A) The 3D structure of FVIII protein. The red color represents the coagulation FVIII light chain, and the green represents the coagulation FVIII heavy chain. The 3D structure was constructed by the SWISS PDB viewer (https://www.rcsb.org/3d-view/3CDZ/1). B) The protein-protein interaction with FVIII. The data were analyzed by STRING (https://string-db.org). VWF: Von Willebrand factor; F9: Coagulation factor IX; F2: Prothrombin; F10: Coagulation factor X; SERPINC1: Antithrombin-III; PROC: Vitamin K-dependent protein C; LMAN1: Lectin, mannose binding 1; MCFD2: Multiple coagulation factor deficiency protein 2; F11: Coagulation factor XI; SERPIND1: Heparin cofactor 2.
|
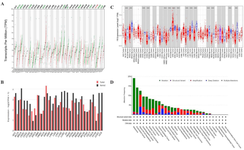
Figure 2. Expression of F8 in different cancers. A) F8 expression profile across all tumor samples and paired normal tissues (dot plot) based on GEPIA2 database. B) F8 expression profile across all tumor samples and paired normal tissues (Bar plot) based on GEPIA2 database. C) F8 expression levels in different tumor types based on TCGA data based on TIMER database.
*p<0.05, **p<0.01, ***p<0.001. D) The gene alterations of the F8 gene in different cancers. Data were downloaded from cBioPortal (https:// www.cbioportal. org/).
|
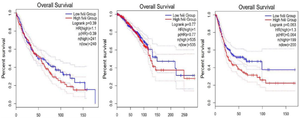

Figure 3. The overall survival curves of patients based on FVIII expression through GEPIA2. A) Lung carcinoma B) Breast cancer C) Bladder cancer.
|
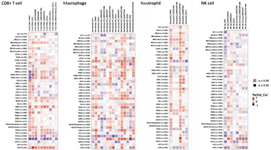

Figure 4. The correlation of F8 expression and tumor immune infiltration. The correlation between F8 gene expression and immune infiltration in 40 types of tumors was analyzed by Timer 2.0 software (http://timer.comp-genomics.org/). The CD8 + T cells, macrophages, neutrophils, and NK cells were selected to assess the correlation with F8 expression in tumors by Spearman’s test. Red color demonstrates a significantly positive correlation (p<0.05), and blue color demonstrates a significantly negative correlation (p<0.05). The color depth shows the value of the correlation coefficient.
|
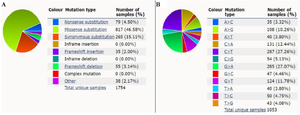
Figure 5. The distribution of different types of mutations for F8 based on COSMIC database (https://cancer.sanger.ac.uk/cosmic). A) A summary of the types of mutation that have been observed in various samples for F8 gene. B) A breakdown of the observed substitution mutations.
|
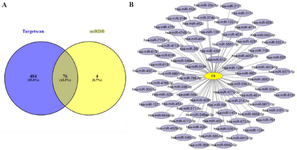
Figure 6. Target miRNA prediction for F8 gene. A) Prediction of a total of 76 common miRNAs that target F8 gene in the two databases miRDB, and TargetScan. B) 76 common F8 gene target miRNAs. The interaction network was constructed by Cytoscape software.
|

Table 1. The expression changes and roles of FVIII in human diseases
|
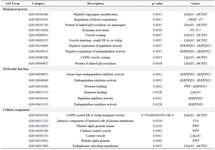
Table 2. Gene ontology enrichment analysis of F8 interaction network based on Enrichr database
|
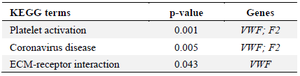
Table 3. The top significant KEGG terms for the F8 interaction network based on Enrichr database
|
|