Effects of Dental Pulp Stem Cell Preconditioning on Osteogenesis using Conditioned Media of Probiotics Bacteria
-
Amini, Fatemeh
-
School of Dentistry, Shahed University of Medical Sciences, Tehran, Iran
-
Bakhtiari, Ronak
-
Department of Pathobiology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
-
 Tabatabaei Ghomsheh , Elham
Department of Restorative Dentistry, School of Dentistry, Shahed University, Tehran, Iran, Tel: +98 9126237114; E-mail: etabatabai@gmail.com
Tabatabaei Ghomsheh , Elham
Department of Restorative Dentistry, School of Dentistry, Shahed University, Tehran, Iran, Tel: +98 9126237114; E-mail: etabatabai@gmail.com
Abstract: Background: Stem cells are used to treat numerous diseases; however, their lifespan is rather short. Factors such as probiotics affect and improve various cell lineage efficacies. The aim of this study was to investigate the effects of probiotics-conditioned media on dental pulp stem cell potentials in osteogenesis.
Methods: The experiment was initiated by culturing Lactobacillus casei and Lactobacillus acidophilus probiotics as well as DPS-7 cells. Bacterial supernatants were separated and concentrated as the conditioned media. The DPS-7 cells were treated with various concentrations of the conditioned media. Furthermore, MTT assay and alkaline phosphatase activity were used. The mRNA expression of three genes (bFGF, EGF-β and BMP-2) involved in osteogenesis was analyzed using a real-time polymerase chain reaction.
Results: The response of dental pulp stem cells to probiotics preconditioning promoted cell proliferation, increased alkaline phosphatase activity and upregulated bFGF and BMP-2 gene expression. Increased expression was significant for BMP-2 and moderate for bFGF; however, it was non-significant for EGF-β. The use of the two probiotics was the most effective.
Conclusion: In general, synergism of the combined probiotics preconditioning induces differentiation of DPS-7 cells into osteoblasts most effectively.
Introduction :
Dental tissues provide accessible sources for stem cells and are easily extracted and processed from exfoliating/extracting teeth. Despite high similarities to Bone Marrow Mesenchymal Stem Cells (BMSCs), Dental Pulp Stem Cells (DPSCs) proliferate faster. The DPSCs can differentiate into adipocytes, neural cells, pulp/dentin tissues and osteocytes 1. Therefore, DPSCs can regenerate damaged or lost teeth. In addition to DPSC advantages, disadvantages such as the lack of cost-effective and safe isolation techniques and short-lived cells with high-regenerative potentials are still controversial. Researchers have developed novel strategies for altering microenvironments of the stem cells and following up their in vitro behaviors. Strategies such as seeding density, coating surfaces, three-dimensional (3D) scaffolds, preconditioning, optimization of cell culture conditions, genetic modifications and hypoxic treatments have improved stem cell functions and characteristics in vitro and in vivo 2. Dental caries is the most common chronic disease that affects teeth function with a high prevalence worldwide. Tooth decay is commonly caused by acid production of numerous micro-organisms in the mouth. The use of probiotic bacteria is an alternative therapy for micro-organism-mediated diseases 3.
Improvement of the efficiency of the stem cells by preconditioning can be achieved using cytokines, hypoxia, pharmacological agents and trophic factors as well as chemical and physical factors 4. First, cells are exposed to the highlighted factors to stimulate cell survival signaling and tolerate the rigorous microenvironment of the transplantation 5. Probiotics with antagonistic activities against pathogens protect against bone damage, regulate inflammatory pathways and upgrade antioxidant qualities 6. In the current study, probiotic preconditioning with two Lactobacillus species was used to increase DPS-7 cell effectiveness. The aim of the current study was to assess the effects of Lactobacillus casei (L. casei) and Lactobacillus acidophilus (L. acidophilus) probiotic preconditioning on DPS-7 cell-line proliferation, osteogenesis and growth factor expression in addition to the synergistic effects of probiotics in these procedures.
Materials and Methods :
Culture of probiotics bacteria: Standard strains of L. casei ATCC 39392 and L. acidophilus ATCC 4356 were provided by the Iranian National Center for Genetic and Biologic Resources, Iranian Biological Resource Center (IBRC), Tehran, Iran. First, probiotic bacteria were recovered on De Man, Rogosa and Sharpe (MRS) agar and then were enriched in liquid MRS media and incubated at 37°C for 2 days.
Preparation of the cell-free supernatants: After preparing the bacterial suspensions, the bacteria were centrifuged at 4000 rpm for 15 min and the supernatants were collected. Then, the two supernatants passed through 0.2-µ membrane filters concentrated using filters of 3 kD (Amicon Ultra‐PL 3; Millipore, USA) and stored as Conditioned Media (CM). The concentration of the CMs was assessed using NanoDrop ND-2000 Spectrophotometer (Thermo Fisher Scientific, USA) at 280 nm 7.
DPS-7 cell culture: Human adult Dental Pulp Stem cells 7 (DPS-7) (IBRC C10371) were provided by the Iranian National Center for Genetic Resources, Iranian Biological Resource Center (IBRC), Tehran, Iran. First, DPS-7 cells were cultured in α-MEM media enriched with 20% of Fetal Bovine Serum (FBS), 2 mM of glutamine and 10 Uml-1 of penicillin/streptomycin. Then, cultured cells were incubated at 37°C with 5% CO2 in humidity.
Treatment of DPS-7 with probiotic-conditioned media: Approximately 3×103 cells were seeded into a 96-well plate and exposed to various doses (0.1-50 µM) of the concentrated probiotic metabolites for 48 hr. The optimal doses of the probiotics with smaller values of the IC50 lethal effects on the cells were assessed using an MTT assay. Briefly, 20 µl of 3(4,5-dimethylthiazol-2-yl)2,5-diphenyltetrazolium bromide (MTT) (Sigma, USA) solution (0.5 mg ml-1) were added to each well and incubated at 37°C for 3 hr. Following the media removal, 100 µl of dimethyl sulfoxide (DMSO) (Sigma, USA) was added to each well and then absorbance was measured at 570 nm using xMark Microplate Reader (Bio-Rad, USA) 7.
Quantification of the growth factor genes expression in DPS-7 cells
The RNA Extraction, reverse transcription and quantitative real-time Polymerase Chain Reaction (qPCR): The fold change method was used for qPCR assessment. Total RNA was extracted from untreated DPS-7 cells using the bioZOL-G RNA Isolation Kit (bioWORLD, USA). Briefly, 500 µl of biozone was added to the cells and continued by phenol/chloroform extraction according to the manufacturer’s instructions. The extracted RNA quality was assessed spectrophotometrically at 260/280 nm using NanoDrop ND-2000 Spectrophotometer (Thermo Fisher Scientific, USA). Complementary DNA (cDNA) was amplified from RNA using commercial kits (Takara, Japan).
Designing primers: Primers of the genes involved in tooth restoration and growth (EGF-β, bFGF and BMP-2) were designed using Allele ID Software (PREMIER Biosoft, USA). In this study, β-actin was used as a reference gene. Sequences of the primers were as follows: F-β-actin, 5¢‑TCAACACCCCAGCCATGTAC-3¢ and R- β-actin, 5¢‑AGTCCATCACGATGCCAGTG-3¢; F-EGF- 5¢‑CC AAACGCCGAAGACTTATCC‑3¢ and R-EGF- 5¢-CTTATTACCGATGGGATAGCCC-3¢; F-bFGF-2 5¢‑ ATGGCAGCCGGGAGCATCACCCACG-3¢ and R-bFGF-2, 5¢‑CAGCTCTTCGCAGACATTGGAAG-3¢; and F-BMP-2, 5′‑TCATAAAACCTGCAACAGCCA ACTCG-3′ and R-BMP-2 5′‑GCTGTACTAGCGAC ACCCAC-3′.
Real-time PCR: Corbett Rotor-Gene TM 3000 (Corbett, Australia) was used for the real-time polymerase chain reaction (real-time PCR) of the genes. Predenaturation of cDNA was carried out at 94°C for 15 min. Then, 40 cycles, including denaturation at 94°C for 20 s, annealing at 58°C for 45 s and extension at 72°C for 10 s were carried out. Each reaction of 25 μl consisted of 2×SYBR Green Mix (Qiagen, Germany), 1 μM of the mixed primers and 0.1 μg of the cDNA. Analyses of amplification curves, melting curves and cycle threshold values (Ct values) were carried out by Rotor-Gene Software (Qiagen, Germany). Then, relevant curves were plotted by GraphPad Prism 5 Software (Dotmatics, USA).
Differentiation of cells into osteocytes and adipocytes: Naturally, DPS-7 cells could differentiate into osteoblasts, adipocytes and odontoblasts in vitro when exposed to specific stimuli. First, 5,000 cells were cultured for the negative control and 15,000 cells in each well of a 6-well plate. After cells reached their 90% confluency, media were replaced with differentiation-inducing media. An osteogenesis differentiation kit containing base and complementary media (StemPro A1007201, Life Technologies, USA) was used to differentiate cells into bones and an adipogenesis differentiation kit (StemPro A1007001, Life Technologies, USA) was used to differentiate cells into adipocytes. Cells were respectively stained with alizarin red S and oil red O and then studied using a light microscope.
Alkaline phosphatase activity assay: Alkaline Phosphatase (ALP) activity is a biochemical marker for osteoblast activity. Briefly, ALP production by the DPS-7 cells was assessed on days 7 and 14 of cell treatment with probiotic CM. First, 105 hDPSC were cultured per well of a 24-well plate. After 7 and 14 days, cells were washed 3 times using Phosphate-Buffered Saline (PBS) solution. Cells were homogenized in 0.5 ml of RIPA buffer (pH=7.5, 0.05% Triton X-100, 1 mM MgCl2, 10 mM Tris-HCl) using sonication and then centrifuged at 12,000 rpm for 10 min at 4°C. Cell degradation contents (0.1 ml of the floating contents) were mixed with 0.5 ml of p-Nitrophenol Phosphate (PNP) (Sigma-Aldrich, USA) solution and 0.5 ml of alkaline buffer solution (Sigma-Aldrich, USA). After incubation at 37°C for 15 min, 1 ml of 0.02 N NaOH was added to the mixture to stop the interaction and adsorption was measured at 405 nm using a spectrophotometer (Bio-Rad, USA) 8. A standard curve of known PNP concentrations was plotted and used to calculate the concentration of the samples. The ALP activity was calculated as the ratio of the PNP nanomoles converted per min to milligrams of the total protein. Total protein was measured using the Bradford assay.
Statistical analysis: GraphPad Prism 5 Software (Dotmatics, USA) was used for data analysis. Moreover, one-way ANOVA and Tukey post hoc tests were used as well. For real-time PCR data analysis, the ΔCt gene was estimated using differences of the corresponding Ct gene, Mann Whitney U test for gene expression in the two distinct groups and a t-test. Values were recorded as mean ±SD (standard deviation) and differences less than 0.05 were reported as significant.
Results :
Preparation of the cell-free supernatants: Two collected supernatants of L. casei and L. acidophilus were filtered and concentrated. The CM concentrations included 7079 and 6860 mg ml-1, respectively (Figure 1).
Morphology and multilineage differentiation capacity of the DPS-7 cells: The DPS-7 cells were cultured and passaged in two-dimensional (2D) culture media. The use of an inverted phase-contrast microscope demonstrated the morphology of the DPS-7 cells at Passage 5 (Figure 2). All treated and untreated groups of DPS-7 cells showed long spindle-shaped and fibroblast-like morphologies. It was verified that treatment with probiotics did not affect the cell morphology and cells were still uniform and smooth with no significant statistical differences in morphologies between the four groups. Culturing DPS-7 cells in osteogenic and adipogenic induction media verified their multilineage functional differentiation capacities. After three weeks, alizarin red S stained calcium deposition and oil red O stained lipid droplets. However, staining intensity was low in all groups following the induction of differentiation. Staining the test groups, the treated groups were a little more chromatic, compared to the control. The plastic adherence characteristics, fibroblastic morphologies and differentiation capacities revealed that the DPS-7 cells were similar to mesenchymal stem cells.
The DPS-7 cell viability assay: The DPS-7 cell viability was investigated after treatment with probiotics alone or in combination using an MTT assay. Cell proliferation increased in a dose-dependent manner when increasing the concentration of the bacterial metabolites. Cell viability was significantly unaffected by the probiotic treatment at concentrations less than 2 µM. Data for Lc-La-DPS-7 showed the highest cell proliferation within the three groups. No statistical differences were seen in the viability of the cells between the Lc-DPS and La-DPS. The two groups revealed almost identical effects on DPS-7. Furthermore, Lc and La concentrations at 5 µM significantly increased DPS-7 cell proliferation (p≤0.05). However, up to 2 µM of Lc and La included significant effects on increasing DPS-7 cell proliferation (p≤0.001) (Figure 3).
Assessment of EGF-β, bFGF and BMP-4 genes expression levels in DPS-7 cells: In this step, primarily extracted RNA concentrations were measured for the four cell groups using similar methods. The RNA concentrations for DPS-7, Lc-DPS-7, La-DPS-7 and Lc‑La-DPS-7 cells included 1292.5, 1364.2, 1102.9 and 1264.1 ng µl-1, respectively (Figure 4).
Real-time PCR results: The BMP-2, bFGF and EGF-β gene products are growth factors affecting bone regeneration and fracture repair. Relative expression levels of these genes in treated and untreated DPS-7 cells were assessed using real-time PCR. Values were normalized to β-actin gene expression. As illustrated in figure 5, significant upregulations of bFGF (p<0.001), EGF-β (p<0.05) and BMP-2 (p<0.001) gene expression were observed after treatment with a combination of probiotics, compared to the untreated group. However, increased levels of EGF-β gene expression were not significant in cells treated with each probiotic alone. The most significant increase belonged to BMP-2 (p<0.001) gene expression in the cell group treated with a combination of probiotics. No significant differences were seen in the three gene expression levels between the Lc-DPS-7 and La-DPS-7. Whereas the gene expression levels of bFGF (p<0.05) and BMP-2 (p<0.01) were significantly higher in the two single-probiotic-treated cells, compared to the untreated cells. Furthermore, bFGF and BMP-2 gene expression levels showed significant increases in Lc-La-DPS-7 cells, compared to the single-probiotic treated cells (p<0.001). Therefore, it could be concluded that a combination of La and Lc probiotics included the highest expression levels of bFGF and BMP-2 genes as potential osteoinductive factors.
Alkaline phosphatase activity: During osteogenic differentiation, cells in the mineralized media produced ALP as an osteoblast marker 9. In the osteogenesis process, differences in increased ALP activity of the four cell groups were significantly time-dependent. On days 7 and 14 of osteogenic induction, the Lc-La-DPS-7 cells showed the most increased ALP activity followed by La‑DPS-7 and Lc-DPS-7 cells, while DPS-7 cells demonstrated the lowest ALP activity. The ALP activity of the cells increased time-dependently between the groups, especially Lc‑La-DPS-7 (p≤0.001), La-DPS-7 (p≤0.05) and Lc-DPS-7 (p≤0.05), compared to DPS-7 on Day 7. After 14 days, differences in ALP activities between the Lc-La-DPS-7 (p≤0.001), La-DPS-7 (p≤0.001) and Lc-DPS-7 (p≤0.001) cells were significant. While no significant differences were reported in the ALP activity of the La-DPS-7 and Lc-DPS-7 cells, treatment of DPS-7 cells with the two probiotics significantly increased ALP activity, compared to that of treatments with each probiotic alone (p≤0.01) (Figure 6).
Discussion :
Recently, stem cell investigations have developed dramatically. Relatively, DPSCs have been popular for their multipotent differentiation and broad therapeutic potential in regenerative medicine and dentistry 10. The benefits and simplified use of these multipotent stem cells in allogeneic stem cell transplantation are due to their immunosuppressive capabilities and low immunogenicity. Nevertheless, the clinical use of DPSCs is restricted by their limited lifespan during proliferation and differentiation. It seems that the broad use of preconditioning strategies can remove this limitation 7,11. Although DPSCs share similar characteristics with stem cells, they are further proliferative. Based on the literature, DPS-7 reveals osteogenic and neurogenic differentiation potentials 12. Growth factor secretion profiles of DPSCs vary with the cell culture conditions, resulting in the alteration of regenerative potentials 13,14. Dominant therapeutic effects of DPSCs depend on the presence of growth factors, cytokines and osteoinductive agents. Possibly, there are several uninvestigated approaches through the modulation of growth factors and cytokine productions 15. For example, researchers’ DPSC treatments with cobalt-chromium and hypoxic induction have led to growth factor alterations in the cells, enhancing their regenerative potentials 14. Studies have reported inductive effects of Lactic Acid Bacteria (LAB) on stem cell osteogenic and adipogenic differentiations 16,17. Furthermore, LAB includes anti-inflammatory and antioxidant activities with no side effects on the stem cells 12,18.
Of the most common markers of osteogenesis, ALP designates the osteogenic differentiation ratio 19. The ALP with prominent roles in biomineralization activities omits the phosphate groups and hence calcification occurs 20-22. The present study assessed ALP on days 7 and 14 of DPSC osteoinduction. In fact, ALP increased between the groups in time-dependent and synergistic manners. This increase was significant for Lc‑La-DPS-7 (p≤0.001), La-DPS-7 (p≤0.05) and Lc-DPS-7 (p≤0.05) cells, compared to DPS-7 cells on day 7 and continued until day 14. Results revealed the synergic effects of the two probiotics on the osteogenic differentiation of DPSCs. Xu et al investigated MSC preconditioning, verifying that Strontium (Sr) folic acid derivate could significantly upregulate the ALP activity of MSCs. Enhancement of the mineralization demonstrated that Sr folic acid promoted MSC osteogenic differentiation 23. Similarly, DPS-7 cells were treated with the two probiotics for 7-14 days in the current study, which showed significant increases in ALP activity, compared to the use of probiotics singularly (p≤0.01). These results verify the synergistic effects of the two probiotics.
Technically, expression markers such as osteogenic differentiation-specific genes can assess osteoblast proliferation and differentiation 24. Cell interaction with the microenvironment regulates the typical differentiation procedure and sustains stable expression of the differentiation-specific genes 25. Morphogenic signals and progenitor/stem cells are the key elements for dental tissue engineering. The BMPs and FGFs families are the major morphogenetic signaling 26. The BMPs, as members of the TGF-β superfamily, are involved in osteoblast differentiation with pleiotropic functions 27. From the family, BMP2 stimulates DPSCs and induces reparative dentinogenesis 28. During DPS-7 development in this study, bFGF, EGF-β and BMP-2 genes were selected to assess DPS-7 differentiation capacity. The potent modulators, bFGF and BMP-2, stimulate cell proliferation and induce osteogenic and neurogenic differentiation. The expression of bFGF and BMP-2 mRNAs was significantly higher in probiotics-induced DPS-7 cells than in untreated cells. Moreover, EGF levels did not statistically differ between the four groups.
Based on the literature, bFGF and BMP-2 enhance the proliferation and mineralization characteristics of DPSCs through various mechanisms. The bFGF stimulates stem-cell proliferation, which begins the cascade of fracture repair mechanisms, including RAS-MAPK, PI3K-AKT, PLCγ, and JAK/STAT signaling pathways. Furthermore, BMP-2 contributes to the morphogenetic signaling pathways through Smad and non-Smad pathways, while EGF-β contributes to the EGFR signaling pathway. The more BMP-2 is expressed, the more osteogenic differentiation is enhanced. These results were similar to those by Toth et al, who suggested that BMP-2 expression induced differentiation of DPSCs into early pre-osteoblasts 29. In this investigation, BMP-2 was expressed more than 5-folds under osteoinductive conditions of the DPSCs. An explanation for the higher BMP-2 gene expression is possibly associated with its contribution to the morphogenetic signaling pathways through Smad and non-Smad pathways. Therefore, it is suggested that the probiotics induced the expression of BMP-2 in DPS-7 cells. Increased production of BMP-2 led to self-induction and subsequent positive self-regulation. In fact, BMPs induced the expression of ALP, type I collagen and osteocalcin 30. Congestion of these factors promoted the proliferation, biomineralization and differentiation of DPS-7 cells into osteoblasts.
Based on the literature, bFGF is involved in cell signaling and maintenance, growth and repair of the tissues. It sustains stemness and regulates osteogenic differentiation and mineralization in stem cells 31. Similar to BMPs, bFGF was actively expressed in DPSCs after preconditioning with the probiotic CM and induction of osteogenic differentiation. Therefore, mRNA transcripts increased significantly, compared to non-preconditioned controls 32. The current results clearly refer to the roles of these genes in controlling and regulating the process of DPSC proliferation and differentiation. The genes functionally contribute to determining cellular characteristics of the probiotic-induced DPSCs. Differences in transcript expression may describe their diverse roles in these processes 32.
In fact, EGF-β contributes to the EGFR signaling pathway, a critical bone regulator necessary for osteogenic differentiation. Based on the studies, EGFR signaling suppresses osteogenesis by downregulating mTOR signaling during osteoblast differentiation 33. The EGFR stops differentiation via trapping osteoprogenitor cells and downregulating transcription factors (Runx2 and Osterix) 34. Moreover, EGF-β is the antagonist of TGF‑β 35. When DPS-7 cells overexpress TGF-β, the biochemical competes with EGF-β for linking to EGFR; hence, EGFR signaling is suppressed and osteogenesis occurs. For this reason, increased EGF-β expression in the current DPS-7 cells was not significant. These results have been supported by Park, Zhu and Linder 33-35. The present results have verified that DPS-7 cells treatment with probiotics improves osteogenic mineralization during cell differentiation. Moreover, combined treatment of DPSCs has indicated the best improvement due to the synergistic effects of the two probiotics.
Conclusion :
In this study, EGF-β, bFGF and BMP-2 were actively expressed in DPSCs after preconditioning with probiotic-conditioned media as well as significant increases in mRNA. The present results have indicated that the increased expression of genes associated with osteoblast lineage in DPSCs is directly a consequence of osteoblast proliferation, mineralization and bone formation under probiotic preconditioning. However, the most increased expression of the genes has been attributed to a combination of the two probiotics, verifying their synergistic effects. Therefore, this strategy can be addressed for DPSC proliferation and osteogenesis in future studies. Preconditioned cells are practically used for dental repair and regeneration. This can relieve difficulties of teething, pain and surgeries.
Ethical Approval :
The current study was approved by the Ethical Committee of Shahed University (IR.SHAHED.REC. 1400.132).
Acknowledgement :
The authors thank the microbiology laboratory staff.
Conflict of Interest :
The authors declare no conflict of interest.
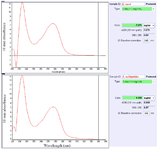
Figure 1. Total protein concentrations of the two cell-free supernatants. Values of the concentrated supernatants included 7079 and 6860 mg ml-1 for Lactobacillus casei and Lactobacillus acidophilus, respectively.
|
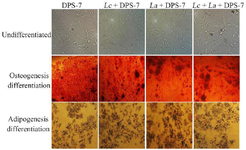
Figure 2. Morphologies and osteogenic/adipogenic differentiation potentials of the dental pulp stem cells. Phase-contrast microscope images of DPS-7 cells at Passage 5 cultured in 2D media showed fibroblastic morphology. The DPS-7 cells (untreated and treated with probiotics) differentiated into osteogenic and adipogenic lineages following three weeks of in vitro differentiation induction culture. Alizarin red S and oil red O staining showed osteogenic and adipogenic differentiations of the DPS-7 cells, respectively. Cell differentiation capacity and morphology revealed that the DPS-7 cells included characteristics similar to that the mesenchymal stem cells did. DPS-7 cells, dental pulp stem cells 7; Lc, Lactobacillus casei; La, Lactobacillus acidophilus.
|
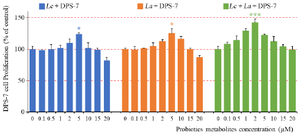
Figure 3. The DPS-7 cell viability after treatment with probiotics. Probiotics improved cell proliferation up to particular concentrations and then decreased it. Each probiotic significantly increased the DPS-7 cell proliferation at 5 µM concentration (p≤0.05). Cell proliferation between the Lc-DPS and La-DPS revealed no statistical differences. However, most DPS-7 cell proliferation significantly increased at 2 µM of Lc and La (p≤0.001). Data is represented as the samples mean and error bars define standard deviations computed from three parallel tests. *p≤0.05; **p≤0.01; ***p≤0.001. DPS-7 cells, dental pulp stem cells 7; Lc, Lactobacillus casei; La, Lactobacillus acidophilus.
|
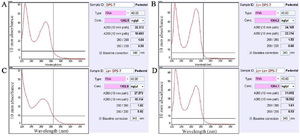

Figure 4. Concentrations of the total extracted RNAs from untreated DPS-7 cells. The four cell-group RNA concentrations included 1292.5, 1364.2, 1102.9 and 1264.1 ng µl-1 for DPS-7, Lc-DPS-7, La-DPS-7 and Lc-La-DPS-7, respectively. DPS-7 cells, dental pulp stem cells 7; Lc, Lactobacillus casei; La, Lactobacillus acidophilus.
|
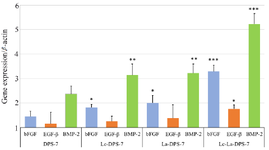
Figure 5. Effects of probiotic treatment on DPS-7 cell gene expression assessed using RT-qPCR. Comparative analysis of bFGF, EGF-β and BMP-2 gene expression levels in DPS-7, Lc-DPS-7, La-DPS-7 and Lc-La-DPS-7 is shown in the figure. The β-actin served as a reference gene. Expression levels of bFGF (p<0.001), EGF-β (p<0.05) and BMP-2 (p<0.001) genes significantly increased in Lc-La-DPS-7, compared to the untreated group rather than single-pro-biotic treated groups.
*p<0.05, **p<0.01, ***p<0.001. DPS-7, dental pulp stem cells 7; Lc, Lactobacillus casei; La, Lactobacillus acidophilus.
|
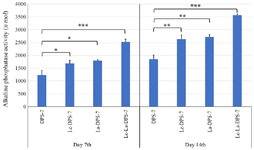
Figure 6. Alkaline phosphatase activity of DPS-7 cells. DPS-7 cells were cultured for 7 and 14 days in osteogenic differentiation media. Effects of DPS-7 preculture with La and Lc probiotics on alkaline phosphatase activity were time-dependent, meaning that differences in alkaline phosphatase activity on day 14 were significantly higher than those on day 7. After 14 days, significant increases were observed in alkaline phosphatase activity of the cells treated with Lc-La (p≤0.001), La (p≤0.05) and Lc (p≤0.05) probiotics, compared to untreated cells. On day 14, differences in alkaline phosphatase activity between the Lc‑La‑DPS-7 (p≤0.001), La-DPS-7 (p≤0.001) and Lc-DPS-7 (p≤0.001) cells were significant while differences in alkaline phosphatase activity of La-DPS-7 and Lc-DPS-7 cells were not significant. The alkaline phosphatase activity of DPS-7 cells treated with the two probiotics was significantly higher than that of cells treated with each probiotic alone (p≤0.01). Data are represented as the samples mean and error bars define standard deviations computed from three parallel tests.
*p≤0.05; **p≤0.01; ***p≤0.001. DPS-7, dental pulp stem cells 7; Lc, Lactobacillus casei; La, Lactobacillus acidophilus.
|
|