MGMT Gene rs1625649 Polymorphism in Iranian Patients with Brain Glioblastoma: A Case Control Study
-
Safaei , Reyhaneh
-
Department of Pathology, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
-
Mojtahedi , Hanieh
-
Molecular Immunology Research Center, Tehran University of Medical Sciences, Tehran, Iran
-
Network of Immunity in Infection, Malignancy and Autoimmunity (NIIMA), Universal Scientific Education and Research Network (USERN), Tehran, Iran
-
Hanaei , Sara
-
Department of Neurosurgery, Imam Khomeini Hospital Complex, Tehran University of Medical Sciences, Tehran, Iran
-
Razavi , Azadehsadat
-
Research Center for Immunodeficiencies, Children's Medical Center, Tehran University of Medical Sciences, Tehran, Iran
-
Esmaeili , Marzie
-
Research Center for Immunodeficiencies, Children's Medical Center, Tehran University of Medical Sciences, Tehran, Iran
-
Sadr, Maryam
-
Molecular Immunology Research Center, Tehran University of Medical Sciences, Tehran, Iran
-
Rezaei, Arezou
-
Research Center for Immunodeficiencies, Children's Medical Center, Tehran University of Medical Sciences, Tehran, Iran
-
Edalatfar, Maryam
-
Department of Neurosurgery, lmam Hossein Hospital, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
Khayat Kashani, Hamidreza
-
Department of Neurosurgery, lmam Hossein Hospital, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
Sadeghi-Naini, Mohsen
-
Department of Neurosurgery, Lorestan University of Medical Sciences, KhorramAbad, Iran
-
Darbeheshti , Farzaneh
-
Department of Genetics, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
-
Gharehdaghi , Jaber
-
Legal Medicine Research Center, Legal Medicine Organization, Tehran, Iran
-
Forouzesh , Mehdi
-
Legal Medicine Research Center, Legal Medicine Organization, Tehran, Iran
-
Ebrahimi , Abdolali
-
Department of Pathology, lmam Hossein Hospital, School of Medicine, Shahid Beheshti University of Medical Sciences, Tehran, Iran
-
 Rezaei, Nima
Research Center for Immunodeficiencies, Children’s Medical Center Hospital, Tehran, Iran, Tel: +98 21 66929234; Fax: +98 21 66929235; E-mail: rezaei_nima@tums.ac.ir
Rezaei, Nima
Research Center for Immunodeficiencies, Children’s Medical Center Hospital, Tehran, Iran, Tel: +98 21 66929234; Fax: +98 21 66929235; E-mail: rezaei_nima@tums.ac.ir
Abstract: Glioblastoma (GBM) is the most common and aggressive primary malignant brain tumor with poor prognosis and high potential of dispersion to other brain tissues in adult. Effective and modern choices of treatment including chemotherapy with alkylating agents marginally extend survival of GBM. However, alkylating agents can lead to highly harmful mismatch during DNA replication causing apoptosis and cell death. Accordingly, O6-Methylguanine-DNA methyltransferase (MGMT) removes alkyl adducts, thereby causing resistance to alkylating drugs. Single-Nucleotide Polymorphisms (SNPs) in MGMT promoter region may play a role in the regulation of MGMT expression and prediction of glioma development risk. In order to evaluate the clinical significance of rs1625649 SNP in the MGMT promoter region of glioblastoma, genomic DNA from a series of 54 patients with GBM and 50 healthy individuals in Iranian population were collected for tetra ARMS PCR amplification. None of the "A" or "C" alleles were associated with tumor occurrence, the "AA" genotype was more frequent in healthy subjects, and the "AC" genotype was 4.6 times more common in patients with GBM. The longest survival time was observed in the "CC" genotype; however, this difference was not statistically significant. On the other hand, homozygous rs1625649 (AA genotype) was significantly associated with a better survival than the cases with heterozygous rs1625649 (CA genotype) or wild type rs1625649 (CC genotype), predicting better response to temozolomide-based chemotherapy.
Introduction :
Glioblastoma (GBM) is the most common and aggressive primary malignant tumor of the central nervous system with poor prognosis and high potential of dispersion to other brain tissues in adults 1. Glioblastomas have a higher prevalence in men and in white people despite their lower incidence in Asians 2,3. The median overall survival for patients diagnosed with GBM is 12-15 months and only 5% of the patients may survive for 5 years after the diagnosis 4. While most GBMs are primary tumors (WHO astrocytoma grade IV), highly invasive, and more commonly occur in older patients, secondary GBMs (grade II) are much less common, associated with more favorable prognostic outcomes, and often occur in younger patients compared to primary ones 5,6. In other words, although primary and secondary GBMs are histologically indistinguishable, they develop from different origins, and are associated with distinct genetic abnormalities caused by molecular alterations 7. Modern choices of treatment for glioblastoma include surgical resection followed by radiation therapy or combined radiation and chemotherapy with alkylating agents such as Temozolomide (TMZ). These conventional therapies marginally increase the average rate of survival, however, despite all the treatment modalities most GBM patients die within 2 years of initial diagnosis 8,9.
Alkylating agents can lead to highly harmful mismatch in O6-methylguanineDNA (O6-MeG) during DNA replication causing apoptosis and cell death 10. O6-Methylguanine DNA methyltransferase (MGMT), as a DNA repair protein, removes alkyl adducts, prevents the formation of cross-links and G>A mutations in the genome, thereby causing resistance to alkylating drugs 11. Epigenetic silencing by aberrant promoter methylation of MGMT, located at chromosome 10q26 , is a clinical predictor to overall survival, since it leads to inefficient repair of DNA alkylation and enhanced response to alkylating agents in GBM 7,12. Both Cytosine phosphate-Guanine (CpG) methylation and Single-Nucleotide Polymorphisms (SNPs) in the MGMT promoter/enhancer region have been found to play important roles in the regulation of MGMT transcription and its downstream protein expression, which can be associated with risk of developing glioma 13,14. In other words, polymorphisms in the MGMT gene may affect the primary structure, expression and DNA repair activity of MGMT 15.
The purpose of this study was to investigate the association of MGMT SNP (rs1625649) in the gene promoter region with GBM and evaluate the clinical relevance of the respective genotypes in Iranian patients with GBMs.
Materials and Methods :
Study population: A series of 54 patients with GBM and 50 healthy individuals with no history of any kind of tumors as control group were selected from Arad and Imam Hossein Hospitals, Tehran, Iran, from 2018 to 2020. The inclusion criteria included patients with newly diagnosed glioblastoma who underwent surgery based on clinical signs (seizures, severe headaches, speech difficulty, and vision disturbance), Magnetic Resonance Imaging (MRI) and histological diagnosis, whereas the samples of control group were obtained from autopsies of healthy subjects.
The most suitable cross section of brain specimens by visual microscopic assessment (>70% neoplastic cells and <50% necrosis) were fixed by Formalin-Fixed Paraffin Embedded (FFPE) protocol. Medical records of all patients were reviewed and follow-up data were collected by accessing patients’ files, and contacting the patients whenever necessary. Basic clinical and demographic data were collected including age, gender, surgical procedure, treatment and overall survival. All investigations described in the current case-control study was approved by the Ethical Committee of Tehran University of Medical Sciences, Tehran, Iran.
DNA extraction: The DNA was isolated from FFPE brain samples by the use of the Phenol: Chloroform Method to extract the amount of DNA required for genotyping, and the purity and yield of DNA were assessed by using a NanoDrop spectrophotometer. Firstly, tissue cores or microdissected tissue were deparaffinized by two-step xylene treatment, which dissolves the paraffin from the tissue, followed by rehydration using a series of descending concentrations with ethanol washes (96, 75, and 50%). Then, all pellets were digested by 20 µl proteinase K (20 mg/ml) (Merck, Germany) and 500 µl lysis buffer [1 M Tris-HCl (pH=8.2), 0.5 M Na2-EDTA, and 100 µl 10% Sodium Dodecyl Sulfate (SDS) (Merck, Germany)]. After overnight incubation at 55°C in a heating block, the mixture of the supernatant with an equal volume of phenol (Merck, Germany) was centrifuged at 12000×g and 4°C for 5 min. Next, the equal volume of phenol/chloroform/isoamyl alcohol mixture (25:24:1) was added to the transformed aqueous layer to separate the proteins from the DNA and then centrifuged for 5 min at 12000´g to separate double-stranded DNA molecules in the aqueous phase from the unwanted proteins and cellular debris. DNA was precipitated by adding 0.1 unit of 3 M sodium acetate (Merck, Germany) and 1 unit of chilled isopropanol (Merck, Germany) and incubation at -20°C for 60 min. After centrifuge for 10 min at 12000xg at 4°C, the dried pellet was washed with 1 ml chilled alcohol 70% twice, to remove contaminations. The DNA was re-suspended in 50 µl of distilled water and stored in the -20°C freezer for further molecular analysis.
Tetra primer amplification refractory mutation system polymerase chain reaction (ARMS PCR): SNP has more applications due to its higher frequency compared with other genetic variations. Tetra primer ARMS PCR is a fast detection method with more simplicity at a negligible cost for SNP genotyping which consists of two sets of primers; the outer and inner primers. The outer primers amplify DNA sequence of interest as the control gene containing SNP, and the largest product of PCR, while the inner primers produce the allele-specific fragments. The sequence of related mutations of MGMT gene with the most pathogenic polymorphisms affecting the glioblastoma was designed using NCBI blast, primer 3 websites and oligo 7 software (Table 1). The tetra ARMS PCR reaction was performed using designed primers and the extracted DNA samples for determining the amplified allele-specific and control fragments. The reaction of tetra primer ARMS PCR was carried out in 20 μl total volume containing 2 ug of template DNA, 0.5 pg of each primer, 5 μl of Taq DNA Polymerase 2x Master Mix RED (Amplicon, Denmark) and 1 μl of DDW, associated with the temperature protocol of; at 95°C for 1 min, further stages in 30 cycles at 95°C for 30 s, 66°C for 50 s, extension at 72°C for 30 s, and a last stage at 72°C for 2 min. In the following, genotype variation from SNPs of interest was visualized by running PCR products in 2% agarose gel electrophoresis.
Statistical analysis: All the experimental data of the allele frequency and genotype of each polymorphism in both groups of patients and controls were statistically evaluated using SPSS 17.0. Qualitative variables were described using frequencies and percentage, and the quantitative ones were described using mean±SD. The association of two qualitative variables were assessed using the Chi-square test, and the survival outcomes in different genotypes were assessed using Kaplan-Meier analysis, with a p-value of <0.05 considered statistically significant in all tests.
Results :
Patient characteristics: The patients were 29 males and 20 females, with mean age of 57.55±13.88 years. The controls were 50 age-gender matched deceased cases, whose samples were obtained from autopsy. The most frequent tumor location was temporal lobe (57.8%), followed by front-al lobe (28.9%), and parietal lobe (8.9%). The other locations were not frequent in the current sample of patients with GBM. While the majority of patients (91.8%) underwent surgical resection as the primary treatment, biopsy was performed in 8.2% of them. As adjuvant therapy, the combination of chemotherapy and radiotherapy was administered for 76.6% of patients, however, in 23.4% of patients, radiotherapy alone was considered as the root of adjuvant therapy. All the patients in the current series had tumor recurrence, and their overall survival time was 12.82±5.74 months. 55.3% of patients had family history of either brain tumor or other cancers (Table 2).
Association of rs1625649 with GBM: Allele and genotype distribution of rs1625649 (C485A) were assessed in the patients and control subjects. The A and C alleles were similarly distributed in patients and controls, with no significant difference (p>0.05). On the other hand, while the AA genotype was significantly more frequent in healthy subjects (p= 0.009), the AC genotype was 4.6 times more frequent in GBM patients (p<0.0005) (Table 3).
Association of rs1625649 with tumor location: In this case tumor location was not influenced by either of rs1625649 genotypes (p=0.9) (Table 4).
Association of rs1625649 genotype and patients’ survival: The mean overall survival was slightly different between three genotypes of rs1625649, with the longest survival time in CC genotype. However, this difference was not statistically different (p=0.77) (Table 5, Figure 1).
Discussion :
The MGMT gene is responsible for encoding a protein with main function of alkyl group removal from O6 position of guanine. Any dysfunction of mis-structure of MGMT may result in accumulation of abnormal MGMT protein in cancer cells, which leads to inefficient activity of alkylating chemotherapeutic agents 16. The SNP of MGMT could possibly alter the gene and/ or protein structure leading to protein malfunction. Importantly, as there could be a heterogeneity in MGMT function in different parts of a tumor in one patient, it could explain the unfavorable responses to treatment despite low levels of MGMT expression 17. The current case-control study was designed to determine the association of MGMT SNP (rs1625649) with GBM in a series of Iranian patients. As explained in detail, while none of the "A" or "C" alleles were associated with tumor occurrence, the "AC" genotype was more frequent in healthy subjects (OR=0.28, p=0.009), and the "CC" genotype was 4.6 times more common in patients with GBM (p=0.0005).
The rs1625649 is located on promotor region of the gene 17, and so far, its association with different cancers was assessed. In a series of 118 patients with GBM, rs1625649 was found in 37% of patients with the "AA" genotype being associated with more favorable progression free survival and lower MGMT protein levels in patients with MGMT-methylated GBM 18. Considering cancers, this SNP was not significantly associated with squamous cell carcinoma of larynx in Chinese population 10. Moreover, it was proposed that some haplotypes of MGMT may influence the sensitivity to alkylating agents more than the SNPs alone 19. It is also reported that rs1625649 was associated with allelic expression imbalance and downregulation of the MGMT promoter activity 20.
Conclusion :
Despite the attempts to prevent biases in the current study, the sample size was not large enough to generalize the results to the whole population of GBM patients. Moreover, and as most of genetic factors are influenced by the environment as well as other genetic variants, the association of SNPs should be evaluated in MGMT-methylated vs. MGMT-unmethylated GBM patients, together with other SNPs. Co-evaluation of serum vs. tumoral levels of proteins could be valuable to determine whether a SNP had effects on protein expression.
Acknowledgement :
This study is supported by a grant from Tehran University of Medical Sciences and Health Services (Grant number: 40721).
Conflict of Interest :
The authors declare no conflict of interest.
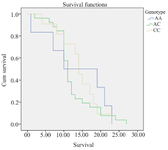

Figure 1. Kaplan Meier Curve for overall survival of GBM patients based on rs1625649 genotype.
|

Table 1. Primers sequences for detecting single nucleotide polymorphism (rs1625649) in MGMT gene
|
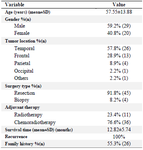
Table 2. Patients' characteristics
|
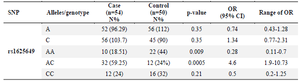
Table 3. Association of rs1625649 with GBM occurrence
|
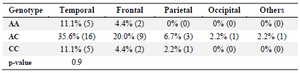
Table 4. Association of tumor location and rs1625649 genotype
|
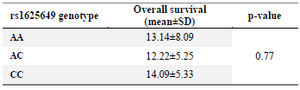
Table 5. Mean overall survival of GBM patients based on rs1625649 genotypes
|
|