CRISPR-Cas System: A Promising Diagnostic Tool for Covid-19
-
 Ebrahimi, Saeedeh
Department of Medical Microbiology (Bacteriology & Virology), Afzalipour Faculty of Medicine, Kerman University of Medical Sciences, Kerman, Iran, E-mail: s_ebrahimi@kmu.ac.ir
Ebrahimi, Saeedeh
Department of Medical Microbiology (Bacteriology & Virology), Afzalipour Faculty of Medicine, Kerman University of Medical Sciences, Kerman, Iran, E-mail: s_ebrahimi@kmu.ac.ir
-
Khanbabaei, Hashem
-
Department of Radiologic Sciences and Medical Physics, Faculty of Allied Medicine, Kerman University of Medical Sciences, Kerman, Iran
-
Fani, Mona
-
Department of Pathobiology and Laboratory Sciences, School of Medicine, North Khorasan University of Medical Sciences, Bojnurd, Bojnurd, Iran
-
Soltani, Saber
-
Department of Virology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
-
Zandi, Milad
-
Department of Virology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
-
Najafimemar, Zahra
-
Infectious Diseases Research Center, Golestan University of Medical Sciences, Gorgan, Iran
Abstract: More than a year has passed since the beginning of the 2019 novel coronavirus diseases (COVID-19) pandemic which has created massive problems globally affecting all aspects of people's life. Due to the emergence of new strains of the SARS-CoV-2, pandemic risk still remains, despite the start of vaccination. Therefore, rapid diagnostic tests are essential to control infection, improve clinical care and stop the spread of the disease. Recently CRISPR-based diagnostic tools have facilitated rapid diagnostic. Here, we review the diagnostic applications of CRISPR-Cas system in COVID-19.
Introduction :
In December 2019, a new beta corona virus called Severe Acute Respiratory Syndrome-Related Coronavirus (SARS-CoV-2) emerged in Wuhan, China and quickly spread throughout the world. After Severe Acute Respiratory Syndrome-Related Coronavirus (SARS-CoV) and Middle East Respiratory Syndrome Coronavirus (MERS-CoV), SARS-CoV-2 is the third zoonotic coronavirus responsible for the pandemic in the current century. SARS-CoV-2 causes the Coronavirus Disease 2019 (COVID-19) 1-4. To date, according to World Health Organization (WHO) reports, more than 100 million people have been infected with COVID-19, and more than 2 million people worldwide have died from the disease. SARS-CoV-2 is a positive+sense RNA virus with a big size genome about ~27 to ~32 kb. SARS-CoV-2 belongs to the Coronaviridae family, which can infect a wide range of birds and mammals such as humans. Due to their broad host range and high frequency recombination in corona viruses, generation of high virulent viruses is common in this family. The most commonly human coronaviruses which can cause a mild infection such as common cold in human includes Human Coronavirus NL63 (HCoV-NL63), Human Coronavirus 229E (HCoV-229E), Human Coronavirus OC43 (HCoV-OC-43) and Human Coronavirus HKU1 (HCoV-HKU1) 1,2,5. Prior to 2003, members of this family were believed to cause only a mild respiratory illness in humans but emergence of Novel coronaviruses SARS-CoV in 2003, MERS-CoV, in 2012 and the most recent SARS-CoV2 in 2019 indicated that they can cause severe acute respiratory syndrome 6. To date several drugs such as Remdesivir, Dexamethasone and inhaled Interferon-beta have been licensed based on the results of randomized controlled trials 7. In addition, recently successes in COVID-19 vaccine development and onset of vaccination against the SARS-CoV-2 have raised hopes for an end to the pandemic. Nonetheless, due to high contagiousness of this disease during the incubation period and the emergence of new strains of the SARS-CoV-2, COVID-19 is still evolving globally and remains a serious health problem 1,8. Therefore, affordable, accurate and rapid methods for detection of COVID-19 is needed to quickly identify infected people and isolate those to avoid further spread of this virus and reduce disease transmission risk.
Various techniques, including RT-qPCR, sequencing-based methods and immunological methods have been used to diagnose SARS-CoV-2. The limitations such as low accuracy and sensitivity of sample preparation, reagents, equipment, and different types of clinical specimens have restricted the common methods of detecting SARS-CoV2 (RT-PCR, Serology); therefore, further research is necessary to find low-cost methods with high sensitivity to detect SARS-CoV2 9-11. In this way, the latest CRISPR-Cas [Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-CRISPR Associated System(CAS)] can be exploited to develop diagnostic or therapeutic approaches. CRISPR-Cas is a part of bacterial acquired immune system in defense against phages and invading genomes. CRISPR-Cas system consist of two parts including CRISPR array and Cas proteins. The CRISPR is an array of short repeated sequences in the genome of bacteria, separated by DNA fragments called spacers. These spacers originate from the phage or invading genomes, which encounter bacteria genome. The Cas proteins are highly diverse and perform many functions including nuclease, helicase, etc. CRISPR array acts as a memory for recognize forging genomes and Cas proteins such as Cas 9 which destroy foreign genomes 12-16. A deeper understanding of the CRISPR-Cas biology can facilitate development of novel diagnostic and therapeutic tools and strategies in the field of infectious diseases.
CRISPR-Cas technology brings promises to diagnose accurately and to treat infectious diseases such as viral and bacterial diseases. In this regard, several studies have shown the potential of CRISPR/Cas technology in the treatment and diagnosis of viral infections (human immunodeficiency virus infection, hepatitis virus infections, etc.) and bacterial infections (Escherichia coli, etc.) 17-20. Rapid and accurate diagnostic tests facilitate early recognition and treatment of infectious diseases. To date, different variants of the CRISPR systems have been introduced for molecular detection of infectious disease. In this review, we have focused on the application of CRISPR-Cas system for COVID-19 diagnosis 21,22.
CRISPR technology for early detection: In recent years, a highly sensitive, precise and rapid technology termed CRISPR-Cas system has caused a revolution in genome editing. Moreover, the CRISPR-Cas system is a well-established genome-editing tool; it has been reported that it may enable rapid detection of the nucleic acid. There are two main classes of CRISPR-Cas systems based on Cas protein: Class 1 (types I/cas3, III/cas10, and IV) and Class 2 CRISPR-Cas system (II/Cas9, V/Cas12, and VI/Cas13) 15,23. Class 1 CRISPR-Cas systems exploit multi-subunit complexes to identify and cleave the target sequence, while Class 2 systems use a single-subunit Cas protein to recognize and cleave the target sequence. Recently, several types of Cas proteins such as Cas9, Cas12, and Cas13 belonging to Class 2 CRISPR-Cas system have been used for detection of nucleic acid. Cas9 and Cas12 endonuclease enzymes are able to cut double-stranded DNA (dsDNA), whereas, Cas13 targets RNA, unlike Cas9 that turns to an inactive state after target cleavage. Nuclease enzyme activity of Cas12 and Cas13 remains active after targeting desired sequences and could cleave both the target and non-target sequences. Therefore, if the target sequence is present in a sample, these enzymes become activated and will start cutting without stopping. Thus, nuclease-mediated degradation of non-target sequences tagged with a fluorophore dye can produce fluorescent signals that can function as an indicator to determine the desired nucleic acids. Therefore, these nucleases can be exploited for nucleic acid detection 15,23-25. To date, several promising approaches have been developed based on CRISPR technology for early diagnosis of COVID-19 such as Specific High-sensitivity Enzymatic Reporter Unlocking (SHERLOCK) system, DNA Endonuclease-Targeted CRISPR Trans Reporter (DETECTR) system, FnCas9 Editor Linked Uniform Detection Assay (FELUDA), Cas3-Operated Nucleic Acid detection (CONAN) and Variant Nucleotide Guard (VaNGuard).
SHERLOCK assay: CRISPR-Cas13a enzyme based Covid-19 detection: Firstly, Feng Zhang and colleagues introduced SHERLOCK for detecting RNA viruses such as of Zika and Dengue viruses. In the present COVID-19 outbreak, the Biosciences Company has received FDA approval for its SHERLOCK-based COVID-19 detection test. Using SHERLOCK method, this company can detect several genes of SARS-CoV-2 within 1 hr 11,26-28. In this method, Cas13-an enzyme that acts as a ribonuclease (RNase) and targeted RNA-instead of DNA is used. Cas13 enzyme becomes activated when binds to RNA targets via specific gRNAs. Activated Cas13 leads to cleavage of RNA target sequences and non-target reporter surrounding ssRNA and results in the production of quantifiable signals 11,26-29. The SHERLOCK detection protocol includes three steps:
1. After extraction of SARS-CoV-2 RNA from patient specimens, the desired sequences such as S or ORF1ab gene of SARS-CoV-2 are converted to cDNA by using RT-RPA or RT-LAMP and subsequently cDNAs are again transcribed to ssRNA using T7 transcription.
2. Then, the CRISPR-Cas13 system complex including Cas13 enzyme, gRNA and fluorescent-labeled nucleotide sequences is added to the amplified ssRNA. After recognition of target sequences via gRNA, Cas13 becomes activated and the fluorescent reporter sequences will be cleaved.
3. Finally, the fluorescent signal produced by cleaving of RNA reporters are revealed (Figure 1).
DETECTR assay
CRISPR-Cas12a Enzyme based Covid-19 detection: Another CRISPR-based detection system for COVID-19 infection called DETECTR technology has been developed by Broughton et al based on CRISPR-Cas 12 30. Firstly, Chen et al established DETECTR technology in 2018 for detecting Human Papillomavirus (HPV) 31. This method is faster than SHERLOCK (~30 min) and unlike to Cas13 used in SHERLOCK Cas12 in DETECTR recognized dsDNA instead of ssRNA 9,15,32. After SARS-CoV-2 RNA extraction from patient specimens, this method includes three steps:
1. The desired sequences of the extracted RNA such as E and N genes of SARS-CoV-2 are reverse transcribed and subsequently isothermal amplified using RT-RPA or RT-LAMP.
2. Then, the cDNA amplicons are directly added to the CRISPR-Cas 12 complex including Cas12 enzyme, gRNA and nucleotide sequences tagged with a fluorescent dye. Cas12 enzyme binds to the target sequence via gRNA; after activation, it will cleave the target sequence and reporter molecules.
3. Lastly, cleavage of reporter molecules by Cas12 generates fluorescence, which can be visualized (Figure 2).
FELUDA assay: CRISPR-Cas9 enzyme based Covid-19 detection: Cas9 enzyme is the most popular genome-editing tool. Cas9 enzyme in combination with a specific gRNA is able to identify and cleave the desired DNA sequence. Several groups have used CRISPR-Cas9 based technology as a diagnostic tool in infectious diseases. Recently Azhar et al has reported FnCas9 enzyme for COVID-19 detection; they named their FnCas9-based method FELUDA assay. FnCas9, an enzyme with has high mismatch sensitivity is found in Francisella novicida 33,34. After extraction of SARS-CoV-2 RNA from patient specimens, this method includes three steps:
1. Extracted RNAs are converted to cDNA by Reverse transcriptase and subsequently amplified using biotin-labelled primers.
2. Then, FnCas9 complex including sgRNA, FAM-labelled tracrRNA, and FnCas9 enzyme are added to the biotin-labelled amplicons. Binding of Ribonucleoprotein (RNP) FnCas9 complex labelled with FAM to the target sequences of positive samples leads to activation of FnCas9 complex, which results in cleavages of the FAM labelled tracrRNA.
3. Finally, adding of gold nanoparticle conjugated with FAM antibody to the previous complex leads to accumulation of antibody-specific binding which consequently makes it further detectable (Figure 3).
CONAN assay : CRISPR- Cas3 enzyme based Covid-19 detection: Yoshimi et al has developed a type of collateral cleavage-based method termed Cas3-Operated Nucleic Acid detection (CONAN) lateral flow assay, which facilitates the rapid (within 40 min), low-cost, and sensitive detection of novel coronavirus SARS-CoV-2 11. As mentioned above, the CRISPR-based diagnostic nucleic acid tests mainly belong to Class 2 CRISPR, but CONAN lateral flow assay is based on Cas3 nuclease, which belongs to Class 1 CRISPR. Cas3 endonuclease, can cleave the desired DNA sequences. Unlike the single Cas12a and Cas13 platforms, CONAN needs multiple Cas proteins (Cas3, 5, 6, 7, 8, and 11) 11,23. After extraction of SARS-CoV-2 RNA from patient specimens, this method includes three steps:
1. The desired sequence of N gene is reverse transcribed and amplified by RT-RPA or RT-LAMP.
2. Then synthesized cDNA is added to Cascade-crRNA complex (including Cas3, Cascade proteins, crRNA and ssDNA reporter probe). Cas3 enzyme binds to the target sequence and becomes activated. The activated Cas3 cleaves the target sequence and ssDNA reporter probe.
3. Finally, the cleavage of reporter probe produces fluorescent signals, which is measurable (Figure 4).
VaNGuard assay: CRISPR-Cas12a variant Enzyme based Covid-19 detection: Similar to other RNA viruses, SARS-CoV-2 has high mutation rates. Since SARS-CoV-2 was first identified a year ago, thousands of SARS-CoV-2 variants such as the UK, South African, Brazilian and Indian variants have been identified. These variants are more contagious and makes it more difficult to control pandemic. On the other hand, such mutations are difficult to detect by conventional methods therefore, early detection of these variants are strongly needed 35,36. Recently, a novel diagnostic tool, termed Variant Nucleotide Guard (VaNGuard) has been developed by a team of scientists from Nanyang Technological University (Singapore). VaNGuard is a CRISPR-Cas 12-based diagnostic assay that using engineered, as Cas12a enzyme can detect wildtype and mutated SARS-CoV-2 within 30 min. This enzyme (enAsCas12a), can tolerate single mismatches when used with SARS-CoV-2-targeting gRNAs. In VaNGuard technique, two different gRNAs against the desired SARS-CoV-2 genome (for example S gene) are applied. Two gRNAs are designed to target the similar sequences among various variants of SARS-CoV-2 35,36.
1. The desired SARS-CoV-2 genome for example S gene is reverse transcribed and amplified by RT-LAMP.
2. Then synthesized cDNA is added to Cascade-crRNA complex including: enAsCas12a, two different gRNAs targeting the S gene and molecule tagged with a fluorescent dye). The enAsCas12a enzyme binds to the target sequence and becomes activated. The activated enAsCas12a cleaves the target sequence and molecule tagged with a fluorescent dye.
3. Finally, the cleavage of molecule tagged with a fluorescent dye produces fluorescent signals which is measurable.
Appling two gRNAs (one was perfectly matched and the other had a single mismatch) with the enzyme enAsCas12a ensures that if there is a mutation in one of the target sites, the other crRNA will still work powerfully, and the fluorescence signal can generate (Figure 5).
Conclusion :
One year after the WHO declared the COVID-19 outbreak a global pandemic, despite vaccine rollout, the disease is still raging worldwide. In addition, the emergence of new SARS-CoV-2 strains threatens to make the pandemic far worse. Various studies have shown that COVID-19 is extremely infectious and the majority of infected people (>80%) with SARS-CoV-2 are asymptomatic or have mild illness. These characteristics (highly infectious viruses and asymptomatic patients and emergence of new SARS-CoV-2 strains) enable the disease to spread throughout the world and lead to extensive damages to the health system and the global economy. Therefore, the fast and specific diagnostic methods to identify infected individuals and quarantine them in order to interrupt the transmission cycle of this deadly virus globally is strongly encouraged. CRISPR-Cas system, the most popular genome editing technology heralds a new era in diagnosis of the SARS-CoV-2. Unlike conventional laboratory methods for detecting COVID-19 such as RT-qPCR, Next Generation Sequencing (NGS), which require highly trained technicians and expensive facilitis, and serological tests that recognize antibodies specific to SARS-CoV-2 in later stages of infection (when opportunities to treat and limit disease transmission have passed), CRISPR-Cas based methods have high specificity and sensitivity, without the need of expensive equipment (Table 1). Given the high precision, specificity, portability, and minimal equipment requirements of CRISPR-Cas based methods, they would be ideal for "easy to perform tests" that are extremely important for diagnosis of Covid19 specially in developing countries or even places with greater risk of infection such as airports, ports, and emergency departments. In this review, we summarized CRISPR-based diagnostic methods, including CRISPR-Cas9, CRISPR-Cas12, CRIS-PR-Cas13 and CRISPR-Cas3, in the development of fast, accurate and portable diagnostic tests in COVID-19.
Conflict of Interest :
The authors declare that they have no conflict of interest.
Funding: Not applicable.
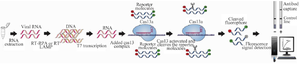
Figure 1. Schematic of CRISPR-based nucleotide detection (SHERLOCK)
|
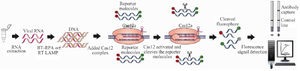
Figure 2. Schematic of CRISPR-based nucleotide detection (DETECTR).
|
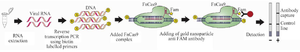
Figure 3. Schematic of CRISPR-based nucleotide detection (FELUDA).
|
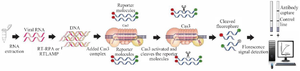
Figure 4. Schematic of CRISPR-based nucleotide detection (CONAN).
|
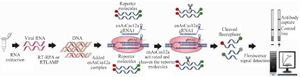
Figure 5. Schematic of CRISPR-based nucleotide detection (VaNGuard).
|
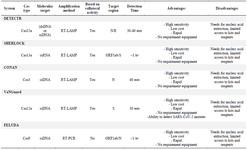
Table 1. Summary of COVID-19 Detection Methods Based on CRISPR/Cas
|
|