An Investigation into the Antifungal Property of Fabaceae using Bioinformatics Tools
-
Arabi, Zahra
-
Drug Design and Bioinformatics Unit, Medical Biotechnology Department, Biotechnology Research Center, Pasteur Institute , Tehran, Iran
-
Faculty of Biological Science, Shahid Beheshti University, Tehran, Iran
-
 Sardari, Soroush
Ph.D., Drug Design and Bioinformatics Unit, Medical Biotechnology Department, Biotechnology Research Center, Pasteur Institute, Tehran, Iran, Tel: +98 21 66405535 Fax: +98 21 66465132 E-mail: ssardari@hotmail.com
Sardari, Soroush
Ph.D., Drug Design and Bioinformatics Unit, Medical Biotechnology Department, Biotechnology Research Center, Pasteur Institute, Tehran, Iran, Tel: +98 21 66405535 Fax: +98 21 66465132 E-mail: ssardari@hotmail.com
-
Drug Design and Bioinformatics Unit, Medical Biotechnology Department, Biotechnology Research Center, Pasteur Institute, Tehran, Iran
Abstract: Chemodiversity in plants provides sources of great value which might be helpful for finding new leads in drug discovery programs. Fabaceae as the third largest family of flowering plants was chosen to investigate its possible antifungal activity. In order to increase the effectiveness of the result, molecular similarity methods and chemical data were used. Twelve plants were selected from Fabaceae and collected from the North and South of Iran. Percolation method with 80% ethanol was used for extraction of collected plants. Antifungal activities of these extracts were determined using broth microdilution method against Candida albicans (C. albicans) ATCC 10231, Aspergillus fumigatus (A. fumigatus) AF 293 and Asperigillus niger (A. niger) ATCC 16404. Extracts with promising activity were screened for toxicity with larvae of Artemia salina (brine shrimp). Dalbergia sissoo, Lathyrus pratensis, Oreophysa microphyalla, Astragalus stepporum, Ebenus stellata, Sophora alopecuroides, Ammodendron persicum and Taverniera cuneifolia showed activity against at least one of the microorganisms used in this study. According to the results of our experiment, the extracts of these plants can be used for further investigation in therapeutic research.
Introduction :
There has been an increasing incidence of fungal infections in recent years, largely due to an increase of AIDS-related opportunistic fungal pathogens and the emergence of resistance strains (1,2). Despite several available antimycotic drugs, the treatment of immuno-compromised patients is still limited due to a number of factors. These include low drug potency, poor solubility of drugs, emergence of resistant strains and drug toxicity.
This situation, coupled with the undesirable side effects of certain antibiotics and the emergence of previously uncommon infections is a serious medical problem. It sounds essential to find new sources of antifungal agents.
Plants are invaluable sources of pharmaceutical products that have drawn the attention of many scientists (3). Fabaceae is the third largest family of angiosperm plant with approximately 730 genera and over 19400 species worldwide, which includes the plants commonly known as legumes (4). The family includes horticultural varieties and many species harvested as crops and for oils, fiber, fuel, timber, medicines, and chemicals. Several types of alkaloids, non-protein amino acids, amines, flavonoids, isoflavonoids, coumarins, phenylpropanoids, anthraquinones, di-, sesqui- and triterpenes, cyanogenic glycosides, protease inhibitors and lectins have been described in this family (5). Additionally, in recent years, reports exist on antifungal activity of peptides and proteins isolated from medicinal plants that have mainly concerned species of Fabaceae family (6).
Legumes are able to fix atmospheric nitrogen via symbiotic Rhizobia in root nodules. Thus nitrogen is easily available for secondary metabolism and it is probably not surprising that nitrogen-containing secondary metabolites (alkaloids, non-protein amino acids, cyanogens, protease inhibitors, lectins) are a common theme in legumes.
The therapeutic effect of medicinal plant is related to the presence of alkaloids, terpenoides, glycosides, phenolic compounds, polysaccharides (7,8). Fabaceae, containing several compounds of these types, can serve as a possible source for new therapeutic agents. Wide variety of species and their compounds and high cost of experimental techniques forced scientists using strategies to reduce the number of plants.
Important criteria which were previously used include Chemotaxonomy: When an interesting lead is found, and either a new (richer) source of the compound or related structures are sought, chemotaxonomy can point to related plant species to screen (9), Traditional use: Many products have traditional uses that are now being investigated to create an evidence base that will facilitate their inclusion in general medical practice (10) and many plant-derived medicines used in traditional medicinal systems have been recorded in pharmacopeias as agents used to treat infections (11), Plant ecological observations: Ecological theories of plant defense can increase the probability of discovering com-pounds with activity in bioassays against human disease targets (12) and Phylogenies: Have great explanatory power and also enable a predictive perspective not offered by previous classifications of plants.
Phylogenetic selection of target species is a new approach to drug discovery in which one strategy could be to select close relative of the most active species for further investigation (13). In order to introduce new methods with higher efficiency, we used chemical data with bioinformatics tools.
Materials and Methods :
Selection strategies
The strategy for selection of cases include informatics based steps and query; the second strategy is application of chemotaxonomy, which are explained in detail below.
1- Cheminformatics strategy
A) Database formation: Literature search was performed to find plants from Fabaceae with antifungal activity. A database was made from the results of the literature search including genera and their antifungal compounds. Non-protein constituents of this database were drawn and converted to SMILES (Simplified Molecular Input Line Entry System) codes using Chem. Draw Ultra (Version 7.01, 2002 Cambridge Soft). The SMILES codes were used for similarity search.
B) Similarity search: Similarity search was performed on SMILES codes of the structures (14); a Tanimoto cutoff score of 0.8 was applied. Following servers were used for similarity search: http://pubchem.ncbi.nlm. nih.gov, http://cactus.nci.nih.gov.
In this way, a subset of compounds similar to antifungal constituents of our database was prepared. These compounds were examined for the presence in Fabaceae as the plant source. Then, the species which existed in flora of Iran were selected.
2- Chemotaxonomy
Since secondary metabolites are often similar within members of a clade, their occurrence or absence might be taken as an indication of common descent and thus relatedness (5).
In our strategy, genera with high antifungal values were chosen from our database. Plant species of tribes from Fabaceae which include these active genera, were selected to achieve either new sources of antifungal compounds or higher concentrations of previously known compounds.
Plant material
Plant materials were collected in March 2007-2008 in the South and August in the North of Iran. The identification of the plants was carried out in our group and for any suspected sample further validation was carried out in Department of Botany of Shahid Beheshti University by Dr Mehrabian.
The voucher specimens were deposited in the herbarium of Pasteur Institute. Aerial parts of plants were air-dried for seven days in shade at room temperature and powdered using an electric grinder. Scientific names of collected plants from Fabaceae family, location and voucher specimen numbers are shown in table 1.
Preparation of extracts
One hundred gm of each plant was extracted using percolation method at room temperature with 300 ml 80% ethanol. This procedure was repeated three times at room temperature away from sunlight. The extracts were evaporated under vacuum at 40 ?C by rotary evaporator (Ika, Germany). The dried residue was subjected to liquid-liquid partition using dichloromethane (DCM, Merck, Germany) and water (3:1) (15). These two fractions were dried in vacuum for further investigation.
Antifungal activity assay
Candida albicans ATCC 10231, Aspergillus niger ATCC 16404, Aspergillus fumigatus AF293 were used as test strains. Plant extracts were dissolved in Dimethyl Sulfoxide (DMSO, Merck, Germany) to prepare the concentration of 20 mg/ml. Broth microdilution was carried out as a standard method for the determination of antifungal activity of plant extracts (16).
Inoculum preparation of yeast
Candida albicans was subcultured on Sabouraud dextrose agar (SDA, Merck, Germany) for 3 days at 35 ?C. The colonies were taken and suspended in sterile saline.
The resulting suspension was adjusted to produce a turbidity of 75-77% of transmittance at 530 nm using spectrophotometer (Bio-Rad, USA). Then the suspension was diluted 1/1000 in Sabouraud maltose broth (SMB, DIFCO, USA) (17).
Inoculum preparation of molds
Each strain of Aspergillus was cultured on Potato dextrose agar (PDA, Merck, Germany) medium for 7 days at 35 ?C. Spores were collected and suspended in a sterile of saline containing Tween 20 (Merck, Germany) (0.1% w/v). Tween 20 was added to facilitate the dispersion of the lipophilic fungal spores in water. After the heavy particles were allowed to set
Result :
Dalbergia sissoo, Sophora alopecuroides, and Lathyrus pratensis were selected using cheminformatic strategy and for the rest chemical data were used. Antifungal activity of ethanolic extracts of these plants was determined. The results of this assay are shown in table 2. The antifungal activity of ethanolic extracts was considered weak. The antifungal assay was repeated using DCM and water phases.
The inhibitory fungal growth by these extracts is summarized in tables 3 and 4. At the same time, control agents were used to compare the effect of extracts. There was no inhibition by DMSO solvent for microorganisms tested at the highest concentration tested (10% V/V).
DCM phase of Dalbergia sissoo showed strong antifungal activity against three microorganisms. Water phase of Lathyrus pratensis, Ebenus stellata, Taverniera cuneifolia and DCM phase of Oreophysa microphyalla, Astragalus stepporum, Sophora alopecuroides, Ammodendron persicum had strong or intermediate activity against C. albicans. Results of toxicity tests which were performed for active phases are shown in table 5.
Discussion :
An emergence of multiple drug resistance in human pathogenic fungi and small number of antifungal classes available stimulated research directed towards the discovery of novel antifungal agents from different sources such as medicinal plants (15). Plants are not only important in traditional medicine, 25% of modern medicines originate from plants first used traditionally (13). Owing to wide variety of plants, over 250,000 species of angiosperms alone (19), it is necessary to develop strategies of selecting plants.
In this paper, we used bioinformatics handling of plant chemical data to come up with a plant selection strategy. In such cheminformatics strategy, antifungal compounds of plant sources with strong antifungal activity were used as queries to find similar structures.
SMILES strings of the structures were applied to search databases of chemical structures. A SMILES string is human understandable, very compact, and if canonicalized, represents a unique string that can be used as a universal identifier for a specific chemical structure. In addition, a chemically correct and comprehensible depiction can be made from any SMILES string symbolizing either a molecule or reaction (20).
Instead of synthesizing and testing similar compounds to find out what effect they have, plant species of Fabaceae which contain similar molecules of the query were selected as new leads for our experiment. Dalbergia sissoo, Lathyrus pratensis and Sophora alopecuroides which were selected by this strategy, showed moderate to strong antifungal bioactivity. This strategy led us to a more reliable way for selection of new target species. It also reduces high cost of experiments and number of plants to be chosen.
There are no previous reports dealing with antifungal activity of plants selected in this study. However, literature is available that discussed the antimicrobial effect of other species of the genera. A herbal preparation containing Dalbergia sissoo and Datura stramoium with cow urine (DSDS), was evaluated for its antibacterial potential against pathogenic strains of gram-positive (Staphylococcus aureus and Streptococcus pneumoniae) and gram-negative (Escherichia coli, Pseudomonas aeruginosa and Klebsiella pneumoniae) bacteria and the result shows that the cow urine extract of DSDS may be used as a potent antiseptic preparation for prevention and treatment of chronic bacterial infections (21). 5,7-Dihydroxy-3 ethylchromone and its 7-O-methyl ether were reported as phytoalexins of Lathyrus odoratus (22) possessing fungitoxicity against Helminthosporium carbonum.
Compounds sophoraflavanone G and kurarinone from the root of Sophora flavescens, were found to have antifungal activity against Streptomyces bikiniensis (23). Finding leads, drug-like compounds that are worthy of further synthetic or biological studies, is a primary goal in a drug discovery project.
Computational search methods have been very useful in this endeavor. Similarity methods are especially useful because little information is needed to formulate a reasonable query. Many implementations of similarity methods are computationally inexpensive, so searching large databases can be routinely performed. The assumption that molecules that are globally similar in structure should exhibit similar biological activity is generally valid (24).
Chemotaxonomy could be another strategy to select species that are likely to contain antifungal compounds. Five out of nine selected species using chemotaxonomy, Astragalus stepporum, Oreophysa microphyalla, Ebenus stellata, Taverniera cuneifolia and Ammodendron persicum, had antifungal activity. Astragalus brachystachys (25) and Astragalus verrucosus (26) are examples of this genus with antimicrobial activity. One study on antimicrobial property of Ebenus haussknechtii by the agar disc diffusion showed that the most biologically active fraction was butanol against bacteria, but it was inactive against Candida albicans (27). Tavernie
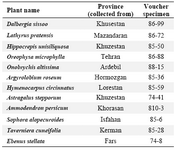
Table 1. Scientific names of collected plants from Fabaceae family, location and voucher specimen numbers
|

Table 2. Antifungal activity of plant ethanolic extracts expressed as Minimum Inhibitory Concentration, MIC (µg/ml)
|

Table 3. Antifungal activity of DCM phase expressed as Minimum Inhibitory Concentration, MIC (µg/ml)
|

Table 4. Antifungal activity of water phase expressed as Minimum Inhibitory Concentration MIC (µg/ml)
|
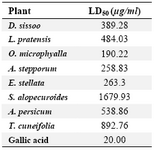
Table 5. The result of toxicity assay on Artemia salina expressed as LD50 (µg/ml)
|
|