An In silico Based Comparison of Drug Interactions in Wild and Mutant Human β-tubulin through Docking Studies
-
 Chellasamy, Selvaakumar
Department of Biotechnology and Bioinformatics, Padmashree Dr.D.Y. Patil University Navi Mumbai, India, Tel: +91 22 27563600; Email: selvaakumarc@gmail.com
Chellasamy, Selvaakumar
Department of Biotechnology and Bioinformatics, Padmashree Dr.D.Y. Patil University Navi Mumbai, India, Tel: +91 22 27563600; Email: selvaakumarc@gmail.com
-
Department of Biotechnology and Bioinformatics, Padmashree Dr.D.Y. Patil University, Navi Mumbai, India, Navi Mumbai, India
Abstract: Background: Tubulin protein being the fundamental unit of microtubules is actively involved in cell division thus making them a potential anti-cancer drug target. In spite of many reported drugs against tubulin, few of them have started developing resistance in human β-tubulin due to amino acid substitutions.
Methods: In this study we generated three mutants (F270V, A364T and Q292E) using Modeller9v10 which were targeted with compounds from higher and lower plants along with marine isolates using iGEMDOCK2.0 to identify their residual interactions.
Results: The mutant F270V does not bring in any increase in the binding affinity in comparison with the taxol-wild type due to their conservative substitutions. However, it increases the volume of the active site. A364T mutant brings a better binding among few of the marine and higher plants isolates due to the substitution of the non-reactive methyl group with the polar residue. But this leads to reduced active site volume. Finally the mutant Q292E from epothilone binding site brings a remarkable change in drug binding in the mutants in comparison with the wild type due to the substitution of uncharged residue with the charged one. But as such there was no change in the volume of the active site observed in them.
Conclusion: Lower plants extracts were reported to exhibit better interactions with the taxol and epothilone binding sites. Whereas marine and higher plants isolates shows significant interactions only in the wild type instead of the mutants. In addition to this, the residual substitutions were also found to alter the conformations of the active sites in mutants.
Introduction :
Tubulin being the fundamental unit of microtubules is critically involved in chromosomal segregation, cell division, motility and intracellular transportation 1. They are made up of α and β subunits alternatively arranged in a lateral and longitudinal manner. The lateral contacts involve the interactions of H1-S2 loop and
helix H3 with the M-loop of the adjacent protofilament. Thus, 13 protofilaments associate laterally and are found to be more electrostatic and less hydrophobic than the longitudinal contacts 2-6. The alternative arrangement of α and β subunits results in longitudinal interactions which are classified into intra- and interdimer interfaces. The intradimeric interface is observed between β and α subunits whereas the interdimericinterface is found between α and β subunits 7-9. Further, the longitudinal contact involves the interaction of H8 of α tubulin with H11-H12, T5, T3 and γ-phosphate of the adjacent subunit. Similarly, T7 of α-tubulin shows interactions with phosphates T2, T1, H7 and Guanine 10.
β-tubulin subunit comprises three distinct domains which include the N-terminal domain, intermediate domain and C-terminal domain. The N-terminal domain harbors the nucleotide and has 6 parallel beta strands (S1-S6) along with the same number of alpha helix (H1-H6). The intermediate domain has strands S7-S10 with three helices H8-H10 which accommodates taxol. The C-terminal is made up of two antiparallel helices H11-H12 which interacts with Microtubule Associated Protein (MAP) 11.
Even though the overall domain architecture remains the same in α- and β-tubulin, still they differentiate themselves both sequentially and structurally. Sequentially they share 40% residual identity, while structurally two positional gaps were observed in β-tubulin at H1-S2 loop (45-46) and S loop (361-368). The larger gap in β-tubulin gets well accommodated by taxol 12. Both the subunits get well associated with GTP; however, hydrolysis is restricted to β-subunit resulting in GTP formation at the exchangeable site. However, it gets sequestered in non-exchangeable site in α subunit 13-23.
Tubulin with its active role in cell division has been considered as a potential anticancer drug target 24. It comprises three distinct drug binding domains which include taxol, vincaalkaloid and colchicine binding sites. Drugs associated with these sites were religiously involved in arresting the mitotic spindle formation 25. Further, taxol and colchicine share overlapping residual interactions towards the inner surface of the microtubules 26. Paclitaxel being the powerful drug for treating several solid tumors including breast, ovarian and non-small cell lung carcinomas 27 prefers to bind with M-loop proximal to S loop resulting in stabilization of lateral interactions of two adjacent protofilaments 28. The important residues associated with taxol binding include V23, D226, H229, T276, R278, R369 and Gly370. These residues were scattered around H1-S2 loop, H7, M-loop and S-loop 20. Apart from paclitaxel, epothilones A and B, eleutherobin and discodermolide have also been reported to bind to the taxol binding site 29.
In spite of all these drug interactions, there is a report of drug resistance among these drugs. The main reason being cited is the residual substitutions associated with the drug resistance. Even though literature supports the correlation between the residual substitutions and drug resistance, but still there is a controversy over their role because of the inclusion of pseudogenes during drug resistance analysis 30. Thus, the debate on the role of residues on drug resistance remains elusive. The reported residual mutations in human β-tubulin include D26E, V60A, S172A, P173A, D197N, E198G, A231T, L240I, F270V, T274P, R282Q, Q292E, R306C, K350N, A364 and Y442C. Several types of resistance have been observed for the drugs like taxol, epothilone, hemiasterlin, 2-methoxyestradiol, vinca alkaloids and indancocine 31-37.
Previous studies report about drugs resistance and the reduced binding affinity of the available drugs, but still not much has been discussed about the drugs targeted against the mutant proteins. In this paper, we tried to target the available chemical compounds from higher plants, lower plants and seaweed secondary metabolites against wild and the mutants of human β-tubulin to investigate their level of interactions.
Materials and Methods :
To begin with, human β-tubulin sequence was downloaded from SwissProt database (accession number: Q9BVA1.1) 38. With no reported crystal structure of human β-tubulin till date, a BLAST-PDB based 39 search was carried out to identify suitable templates. Out of the reported hits, 1JFF was downloaded from Protein Data Bank (www.rcsb.org) 40 which was further considered as a template (1JFF-B chain) for the modeling of the query sequence. Pairwise alignment of the template and the query sequence were generated using Modeller9v10 41 (Figure 1). Using single protocol template from Modeller, 30 structures were generated. Of these generated structures, the model with the least DOPE score (Discrete Optimized Protein Energy) was considered for energy minimization with 100 iterations using steepest descent in SwissPdbViewer 42. The template and the modelled structures were superimposed with each other (Figure 2 A-C). The DOPE score of the template was -49714.55 and the modelled structure was -55870.25. The optimized structure was further validated using PROCHECK of SAVES server (http://nihserver.mbi.ucla.edu/SAVES/) 43. Similar methods were followed for the generation of mutants F270V, A364T and Q292E, respectively.
Conversely, chemical compounds needed for docking against wild and the mutants of human β-tubulin were obtained from marine flora. The chemical compounds were obtained from seaweed secondary metabolite database (www.swmd.co.in) 44. All these available compounds were isolated form red algae Laurencia obtuse (RL) and Galaxaura marginata (RG type). Some of these compounds were reported to be cytotoxic against cancer cell lines 45-49. During our previous study, out of the 517 compounds, we could identify four lead compounds named RG012 (6β, 24ε-Dihydroxycholesta-4, 25-dien-3-one), RL381 (diterpenes containing bromine), RL366 (15-epi-prostaglandin A2 diester), and RL376 (diterpenes containing bromine) which exhibit better binding to wild type human β-tubulin protein 50. All these selected compounds were taken into consideration for docking against the wild and mutant human β-tubulin proteins in the current study.
Next, through literature survey, we identified chemical compounds both from higher and lower plants. To begin with, the compounds from higher plants include Berbamine, Butulinic acid, Camptothecin, Cucurbitacin, Ellipticine, Flavopiridol, Homoharringtonine, Silvestrol, Berberine, Daphnoretin and Podophyllotoxin. Regarding their sources, Berbamine is extracted from Berberis vulgaris with a reported apoptosis in human myeloma cells 51,52. Betulinic acid is a pentacyclic triterpenoid with reported antiretroviral, antimalarial and anti-inflammatory activity and anticancer properties extracted from the bark of Betula
pubescens 53,54.
Camptothecin isolated from the bark and stem of Camptotheca acuminate is a cytotoxic quinoline alkaloid which inhibits the DNA enzyme topoisomerase I 55. Cucurbitacins are from the family of Cucurbitaceae with anti-cancer and anti-inflammatory activities 56. Podophyllotoxin is a non-alkaloid toxin lignan extracted from the roots and rhizomes of Podophyllum species which is again an antitumor agent 57-59. Silvestrol is isolated from Aglaia foveolata and results in apoptosis in the cell lines of hormone-dependent human prostate cancer 60. Homoharringtonine is extracted from Cephalotaxus harringtonia which is identified to be a cytotoxic alkaloid and is generally reported to block the progression of cells from G1 phase in to S phase and G2 phase into M phase 61. Flavopiridol is an indigenous plant from India, which can arrest cell cycle progression at the G1/S and G2/M boundaries 62. Daphnoretin obtained from Wikstroemia indica exhibits strong antiviral and anti-tumor activities with a report of cell cycle arrest in the G2/M phase 63. Ellipticine is isolated from Apocyanaceae plants with established antitumor and anti-HIV activities with their limited toxic side effects and their lack of hematological toxicity 64.
Regarding lower plants, Aclarubicin is produced by Streptomyces galilaeus actively used in the treatment of cancer 65. Daunorubicin and its derivative, doxorubicinare antitumor anthracycline antibiotics are produced by Streptomyces peucetius 66. Blasticidin is a potent antifungal and cytotoxic peptidyl nucleoside antibiotic from Streptomyces griseochromogenes which plays a significant role in controlling prokaryotic and eukaryotic cell growth 67. Chartreusin is a potent antitumor agent with a mixed polyketide-carbohydrate structure produced by Streptomyces chartreusis 68. Neothramycin has been isolated from Streptomyces MC916-C4 which is a potent antitumor antibiotic of the pyrrolo (l, 4) benzodiazepine group 69,70. Pirarubicin is an anthracycline drug which has a diversified antitumor activity 71,72. All these chemical compounds were downloaded from Pubchem database. The chemical structures were obtained from chemical book (www.chemicalbook.com) (Table 1). The summary of the chemical compounds of lower and higher floras along with marine derivates are tabulated in table 2.
All these selected compounds were charged using gasteiger charges available in CHIMERA software 73. Energy was minimized using PRODRG and hydrogen atoms were added to them. Then these compounds were considered for docking using iGEMDOCK software 74. The active sites of taxol (V23, D226, H229, T276, R278,R369 and Gly370) and epothilone (H227, A231, T274, R276, R282 and Q292) 75 were selected for docking (Figure 3). Next, only the residual mutants with deleterious effects were considered for homology modelling. This was identified using amino acid substitution (AAS) tools like Polyphen2 (polymorphism phenotyping) 76,77, PANTHER (protein analysis through evolutionary relationship) 78 and I-Mutant 2.0 79. The list of residual substitutions which bring in deleterious effects are tabulated (Table 3) (for detailed report refer) 80. Out of ten deleterious sites identified, only three were selected as these substitutions were found to be proximal to the drug binding sites. Unfortunately, the rest of the positions were located at distal sites of both taxol and epothilone binding sites. Docking of the lower and higher plants along with marine isolates were carried out using iGEMDOCK for taxol-wild, taxol-mutants (F270V, A364T), epothilone-wild and epothilone-mutant (Q292E). The docking study followed the accurate docking protocol which was very slow with population size of 800 and the number of solution equal to 10. The generation number was maintained at 80. Similar protocol was followed for taxol and epothilone binding pockets. Finally, we measured the size of the normal grooves for wild and the mutants of taxol and epothilone binding sites using SwissPdbViewer.
Results :
In the present study, the chemical compounds obtained from different resources like lower plants, higher plants and marine isolates were docked against the wild and the mutant human β-tubulin proteins. This study was mainly instigated to identify the level of interactions exhibited by both the wild and the mutants for taxol resistance (F270V, A364) and epothilone resistance (Q292). For this study, we considered the results of our previously reported lead molecules from seaweed secondary metabolites. Simultaneously, the chemical compounds from lower and higher plants were also taken into consideration.
First of all, the wild tubulin when docked with these compounds exhibited better binding to Aclarubicin. Next, the compounds from higher plants like Berbamine, Butulinic acid, Cucurbitacin, taxol and Podophyllotoxin exhibited better interactions in the wild type in the taxol binding site similar to the marine compounds. Next, the mutants were considered for docking with all the available compounds. Here, Campothecin, Flavopiridol and Berberine exhibited better interactions in mutant1 (F270V). This site is proximal to M-loop and is also the taxol binding site. With respect to mutant2 (A364T), the chemical compounds like Camptothecin, Ellipticine, Flavopiridol, Berberine, Daphnoretin showed better chemical interactions. In epothilone binding site, lower plant compounds like Aclarubicin and Blasticidin showed better interaction in the wild type. The compounds like Neothramycin, Daunorubicin and Aclarubicin showed slightly better interaction with the mutation of glutamate with Glu at residual position 292. Thus, our overall study confirms that marine compounds exhibit better interaction in this mutant protein except RL376. Among higher plant products, Camptothecin, Ellipticine, Epothilone, Podophyllotoxin exhibit better interaction in the wild type of epothilone binding site. But compounds like Berbamine, Flavopiridol, Berberine and Daphnoretin conveyed better interactions again in the mutant (Table 4). General survey confirms a similar type of interactions exhibited both by wild and the mutant proteins. Thus, a closer inspection of the pocket was much needed for both sets of proteins. Therefore, we measured the groove size for wild, F270V and A364T of the taxol binding site. The volume was 550, 607 and 220 Å3 respectively. However, for epothilone binding pockets, the wild and the mutant (Q292E) maintained same volume of 8 Å3. There was an increase in pocket volume observed for mutant1 in comparison with the wild type due to non-conservative residual substitution. Similarly, the volume of mutant2 decreases in comparison with the wild type in spite of conservative substitution. However, in epothilone binding site, the conservative substitution does not bring in any change in the volume both in wild and the mutant and the grooves were found to be discontinuous with the maximum volume size of 8 Å3 (Figure 4).
Discussion :
During the docking of the wild and the mutants against the available compounds, only three mutants were generated which were proximal to the drug binding site, and were also reported to be lethal. Thus, the generated mutants were christened as mutant1 (F270V) mutant2 (A364T) for the taxol binding site. Similarly, mutant (Q292E) was selected for epothilone binding site. Docked structures were separately investigated for their interactions using CHIMERA software. We observed that Neothramycin exhibits interactions with the taxol binding site through P272 and R276 (Figure 5A). Similar interactions were exhibited by mutant1 (Figure 6A). Also mutant2 exhibited interaction with L361 (Figure 6B). Likewise, RL381 displayed interactions in the wild type of taxol binding site with H227, R282, and R359 (Figure 5B). Camptothecin shows interaction with T274 (Figure 5C) while taxol based mutants, irrespective of their higher binding energy, do not display any interactions. Ellipticine also confirms no significant binding in mutant2. Flavopiridol does not show any significant interactions in the wild taxol binding site. But mutant1 shows interaction with H227 (Figure 6C). Also mutant2 shows interactions with T274 and A275 (Figure 6D). Berberine also shows no significant interaction in the wild type and mutant2 of taxol binding site but mutant1 confirms their interaction with R276 (Figure 6E). Again with Daphnoretin, both wild and mutant2 show no interactions, but mutant1 interacts with R276 (Figure 6F). Next, regarding the epothilone binding site, Neothramycin binds with mutant at T274 (Figure 7A). RL366 interacts in the mutant with R276 (Figure 7B). However, the rest of the compounds like RL381, Flavopiridol, Berberine and Daphnoretin exhibited no significant interactions. Since no remarkable change in the pattern of interactions both in wild and the mutant was observed, we considered the groove analysis for the taxol and the epothilone binding site. This study confirms an increase in the channel size for mutant1 and a decrease in the channel size for mutant2 in comparison with wild type. Moreover, epothilone binding site remains undisturbed with the residual substitution. We hypothesized that the drug resistance especially for taxol binding drugs could be related to the conformational changes in the active site pocket due to residual mutations.
Conclusion :
The residual interaction analysis in wild and the mutants of taxol and epothilone binding sites reveals a better drug binding with the lower plants in the wild types of taxol and epothilone. However, the marine compound and the higher plants in spite of their increase in binding score could not establish a better contact with the residues of both the binding sites. Further, the chemical compounds showed interactions with the charged residues like arginine and histidine for better drug binding.
Acknowledgement :
The authors acknowledge the cooperation of Padmashree Dr. D.Y. Patil University, CBD Belapur for providing the facilities to carry out this work.

Figure 1. Pairwise sequence alignment of template (1JFF-B chain) with the query sequence of human β-tubulin
|
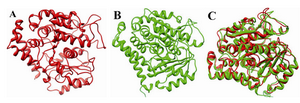
Figure 2. 3D structure of the template; A) the modelled structure of human β-tubulin; B) and its structural superimposition C)
|
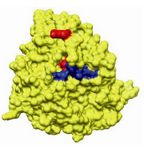
Figure 3. Drug binding sites in human β-tubulin for epo-thilone (Red) and taxol (Blue)
|
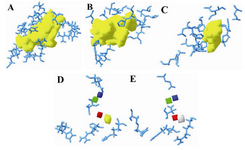
Figure 4. The groove of the active site pockets in wild and the mutant human β-tubulin; A) wild-taxol binding site; B) mutant1-taxol binding site; C) mutant2-taxol binding site; D) wild-epothilone binding site; E) mutant-epothilone binding site
|
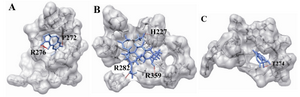
Figure 5. The docking of chemical compounds in the taxol binding sites of wild human β-tubulin; A) Neothramycin-taxol-wild; B) RL381-taxol-wild; C) Camptothecin-taxol-wild
|
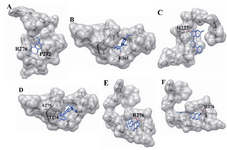
Figure 6. The docking of chemical compounds in the taxol binding sites of mutant human β-tubulin. A) Neothramycin-taxol-mutant1; B) Flavopiridol-taxol-mutant1; C) Berberine-taxol-mutant1; D) Daphnoretin-taxol-mutant1; E) Neothramycin-taxol-mutant2; F) Flavopiridol-taxol-mutant2
|
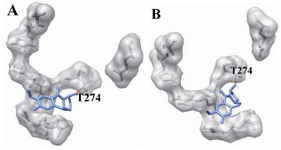
Figure 7. The docking of chemical compounds in the epothilone binding sites of mutant (Q292E) human β-tubulin. A) Neothramycin-epothilone-mutant; B) RL366-epothilone-mutant
|
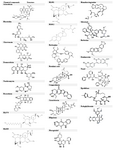
Table 1. Chemical structures of higher and lower plants along with marine compounds
|
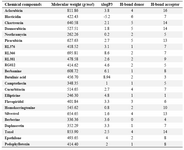
Table 2. Summary of chemical compounds of lower and higher floras along with marine derivatives
|
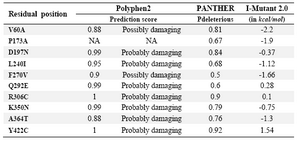
Table 3. Predicting the effect of residual substitutions through amino acid substitutions tools Polyphen2, PANTHER and I-Mutant2.0
|
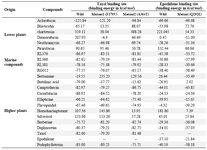
Table 4. Docking of chemical compounds from lower, higher and marine resources against taxol and epothilone binding sites in wild and mutant proteins
|
|